PROOF OF CONCEPT
Integrating Large Language Models Into Clinical Decision Support Systems: A Novel Approach to UTI Diagnosis and Treatments
Manoj Jain, MD1  ; Hiren Pokharna, MD2
; Hiren Pokharna, MD2  ; Sridhar Sunkara, MS3
; Sridhar Sunkara, MS3  ; Sanjeev Bora, MCA4
; Sanjeev Bora, MCA4  ; Kiran Ponamgi, BS5
; Kiran Ponamgi, BS5  ; Rohan Dang Sharma, BS6
; Rohan Dang Sharma, BS6  ; Amar Gupta, PhD7
; Amar Gupta, PhD7 
1Adjunct Professor, Department of Health Policy and Management, Rollins School of Public Health, Emory University, Atlanta Georgia, USA; 2Infectious Disease Physician, Health Science Center, University of Tennessee, Memphis, Tennessee, USA; 3Chief Executive Officer, eBiz Solutions, LLC., Memphis, Tennessee, USA; 4Chief Thinking Officer, eBiz Solutions, LLC., Memphis, Tennessee, USA; 5Project Manager, eBiz Solutions, LLC. , Memphis, Tennessee, USA; 6Research Intern, Computer Science & Artificial Intelligence Laboratory (CSAIL), Massachusetts Institute of Technology, Cambridge, Massachusetts, USA; 7Research Scientist, Computer Science & Artificial Intelligence Laboratory (CSAIL), Massachusetts Institute of Technology, Cambridge, Massachusetts, USA
Keywords: artificial intelligence, clinical decision support systems, diagnosis, large language models, urinary tract infections
Abstract
Background: Urinary tract infections (UTIs) are among the most common bacterial infections globally, leading to significant healthcare expenditures and frequent misdiagnoses. In the United States, UTIs account for approximately 380,600 preventable adult inpatient stays annually, costing $2.55 billion (€ 2.26 billion). Current Clinical Decision Support Systems (CDSS) are often static, lack personalized recommendations, and do not incorporate real-time clinician feedback. AI (alternative)-driven CDSS, leveraging large language models, offers the potential to enhance diagnostic precision, optimize antibiotic use, and improve workflow efficiency.
Methods: We propose automatic photographic apparatus (3RDI), a conceptual framework for an AI-driven CDSS for UTI management, utilizing the novel DETNQ (Diagnosis, Evidence, Treatment Plan, Notes, Quality) structure to organize clinical outputs. This framework envisions a system that would process comprehensive patient data, including medical history, symptoms, laboratory results, and medication records. 3RDI’s design incorporates a day-wise iterative process for continuous feedback, allowing clinicians to refine the system’s recommendations and establish a collaborative human-AI decision-making environment.
Findings: Our conceptual development demonstrates the integration of an adaptable, clinician-driven feedback mechanism within CDSS architecture. Preliminary prototyping suggests the potential for structured patient data presentation using the DETNQ format and adaptability to specific clinical contexts. The proposed framework addresses key limitations in current CDSS, particularly around clinician engagement, workflow integration, and continuous learning capabilities.
Interpretation: The 3RDI conceptual framework offers a promising direction for future CDSS development, particularly for UTI management. Its emphasis on a continuous learning system and clinician feedback provides a blueprint for AI systems that could enhance diagnostic precision while gaining clinician trust. Future work should focus on empirical validation through controlled studies and iterative refinement based on real-world clinical implementation.
Plain Language Summary
Urinary tract infections (UTIs) are one of the most common infections and often lead to misdiagnoses, unnecessary tests, and expensive hospital stays. Current decision-making tools do not update new patient information or feedback from clinicians. This study aimed to develop a conceptual framework to improve UTI diagnosis and treatment.
The research team designed a framework for an AI (artificial intelligence)-powered Clinical Decision Support System—software programs that assist doctors in making clinical decisions. The framework includes a mechanism for the system to learn and improve based on feedback from doctors, ensuring it could adapt to real-world clinical needs. The researchers demonstrated that such a system would be technically feasible to build and could potentially provide valuable information to healthcare providers.
Citation: Telehealth and Medicine Today © 2025, 10: 554
DOI: https://doi.org/10.30953/thmt.v10.554
Copyright: © 2025 The Authors. This is an open-access article distributed in accordance with the Creative Commons Attribution Non-Commercial (CC BY-NC 4.0) license, which permits others to distribute, adapt, enhance this work non-commercially, and license their derivative works on different terms, provided the original work is properly cited and the use is non-commercial. See http://creativecommons.org/licenses/by-nc/4.0. The authors of this article own the copyright.
Received: January 27, 2025; Accepted: May 1, 2025; Published: July 2, 2025
Corresponding Author: Manoj Jain, Email: mjainmd1@gmail.com
Clinical Decision Support Systems (CDSS) are software programs that assist doctors in making clinical decisions and provide support to reduce errors and increase decision-making speed, operating in conjunction with the expertise of medical professionals.1 Advances in artificial intelligence (AI) present an opportunity to improve significantly upon CDSS offerings and are a key element of the latest generation of CDSS.2 The integration of machine learning (ML) algorithms, natural language processing (NLP), deep learning, and large language models (LLMs) enables faster data processing, personalization, and higher accuracy in CDSS.3,4 One potential application of an AI-driven CDSS is for the diagnosis and treatment of urinary tract infections (UTIs).
Urinary tract infections common bacterial infections that affect the urinary system.5 Symptoms include frequent urges to urinate, painful urination, cloudy or foul-smelling urine, and pelvic pain. Diagnosis usually involves analyzing urine samples to detect the presence of bacteria and white blood cells. In clinical microbiology diagnostic laboratories, UTIs are the most common infectious disease tested for.6 As a result, they are responsible for significant expenditure in healthcare systems globally.6 They are also commonly misdiagnosed.7 In 2017, UTIs caused 380,600 preventable adult inpatient stays, which cost the US healthcare system $2.55 billion (€ 2.26 billion). This accounted for 10.8% of all potentially preventable inpatient stays and 14.2% of pediatric inpatient stays.8 Excess expenditure, preventable inpatient stays, and misdiagnoses could potentially be reduced by leveraging AI-based CDSS to diagnose and treat UTIs.9
The Evolution of CDSS and Current State of the Art
The CDSSs have evolved significantly over the past few decades. Early systems were primarily rule based, utilizing “if-then” logic to provide recommendations.10 The second generation incorporated probabilistic and Bayesian approaches, allowing for more nuanced decision support.11 The history of CDSS approaches employing a variety of ML techniques is outlined in Table 1.
Several studies explored the potential of AI and ML models to diagnose medical conditions, some of which focused specifically on UTIs. Current AI-driven CDSS are built using ML algorithms such as neural networks, decision tree algorithms, support vector machines (SVMs), Bayesian networks, and ensemble learning methods.4 These deliver predictive analytics and decision recommendations based on input data. For example, researchers at Cardiff University used a Random Forest Classifier model to diagnose UTI using white blood cell count, bacterial count, red blood cell count, and epithelial cell count and achieved >95% classification sensitivity.9
Similarly, de Vires et al.12 developed a semi-supervised CDSS system for UTI prediction using the RESSEL (reliable semi-supervised ensemble learning method), which outperformed traditional urinalysis and urine cultures to identify UTIs. Their study emphasized that incorporating urinalysis results alongside Gram stain and other readily available parameters allowed clinicians to make better-informed decisions without prematurely resorting to antibiotics.12 This system predicted UTIs more accurately and helped reduce the overuse of antibiotics by holding off prescriptions until culture results were available, thus supporting a key component of antibiotic stewardship.
In addition, CDSSs may also be built using NLP, which enables the analysis of unstructured clinical text, such as clinician notes, discharge summaries, and medical literature.4 Deep learning also allows for more intelligent CDSSs, utilizing convolutional neural networks to extract complex patterns from heterogeneous, multimodal medical data.4 LLMs—one form of AI—have not been as heavily tested as high-quality, open-sourced models, which have only recently been made available. For instance, LLMs have demonstrated potential in improving diagnostic accuracy for common illnesses, as highlighted by Gupta et al., who noted that LLMs like GPT-4 and Gemini show promise for enhancing efficiency and diagnostic precision in healthcare settings.13
One group of researchers developing CDSSs using LLMs implemented retrieval-augmented generation (RAG) techniques.14 This approach, designed to assist pharmacists in addressing drug-related problems (DRPs), utilized the Pharmaceutical Care Network Europe (PCNE) classification to structure its outputs. The RAG framework enabled effective retrieval and contextual integration of pharmaceutical information, incorporating tools to index and retrieve drug-related data for enhanced relevance. The research highlighted the potential of LLMs in CDSS, showing that pharmacists working alongside the RAG-based system could significantly reduce DRPs.14 Liu et al.15 also illustrated how AI-generated recommendations from ChatGPT can enhance clinical decision-making logic, underscoring AI’s role in refining clinician workflows by providing novel perspectives and suggestions that complement human judgment.
The Limitations of Current CDSS Approaches and the Unique Value of AI for UTI Management
Despite these advancements, current AI-driven systems face several significant limitations (Table 2). These factors collectively demonstrate why AI-based CDSS approaches are uniquely positioned to address the challenges in UTI management that other approaches have failed to resolve effectively.
| Limitations | Examples |
| Lack of continuous learning | Most of the machine-learning-enabled clinical decision software on the market today is “locked” software, in which the algorithm, having been trained on an initial data set and tested, does not continuously learn and automatically adapt thereafter.16 This limitation prevents systems from improving through real-world use. |
| Limited clinician feedback integration | Few systems effectively incorporate iterative clinician inputs and continuous feedback loops to drive meaningful improvements.17 This gap diminishes the potential for personalization and adaptation to specific institutional practices. |
| Workflow integration challenges | Many CDSS fail to integrate seamlessly into clinical workflows, creating additional burden rather than reducing cognitive load.18 |
| Transparency and trust issues | The “black-box” nature of many AI models limits clinician trust and adoption, as providers cannot easily understand the reasoning behind recommendations.19 |
| Narrow patient context processing | Many existing models do not yet fully integrate a broad range of patient-specific factors, such as age, medical history, medication, and comorbidities, which are essential for personalized care.20 |
| Challenges | |
| Pattern recognition across multiple data points | A diagnosis of UTI involves synthesizing information from symptoms, laboratory values, patient history, and comorbidities—a task where ML algorithms excel. Recent studies demonstrate that 30–40% of UTI misdiagnoses stem from pattern recognition errors in complex patient presentations. Traditional clinical approaches often struggle with this complexity, particularly in patients with atypical presentations or multiple confounding conditions. |
| Integration of dynamic antibiotic resistance patterns | Selecting appropriate antibiotics requires integrating patient-specific factors with constantly evolving local antibiotic resistance patterns. Traditional clinical pathways have shown limited effectiveness in reducing preventable hospitalizations because they often cannot adapt quickly to changing resistance patterns. The AI systems can continuously update treatment recommendations based on real-time susceptibility data from hospital laboratories, something static guidelines cannot achieve. |
| Cognitive load reduction for high-volume conditions | The high prevalence of UTIs makes them an ideal candidate for AI augmentation, as clinicians face decision fatigue when repeatedly evaluating similar cases. Comparative studies reveal that AI-augmented diagnostic processes for infectious diseases can achieve a 22% reduction in diagnostic errors compared to standard clinical pathways and a 17% improvement compared to non-AI CDSS. |
| Standardization with personalization | Current approaches to UTI management often oscillate between overly rigid protocols that ignore patient specificity or highly variable care that depends heavily on individual clinician knowledge. AI-driven systems can maintain standardization while offering personalized recommendations based on individual patient factors. |
| AI: artificial intelligence; CDSS: Clinical Decision Support System; ML: machine learning; UTI: urinary tract infection. | |
The Proposed Automatic Photographic Apparatus Framework
This article presents “automatic photographic apparatus (3RDI),” a conceptual framework for an LLM-driven CDSS designed to address the limitations identified above and advance the field of clinical decision support for UTI diagnosis and management. The 3RDI framework focuses on several key innovations. These include (1) a day-wise iterative process that aligns with clinical workflows and allows both practitioners and the AI system to learn and adapt over time, (2) a novel DETNQ (Diagnosis, Evidence, Treatment Plan, Notes, Quality) structure to organize clinical information, a continuous learning system (CLS) that incorporates clinician feedback to improve recommendations, and (3) the integration of a broad range of patient data, including demographics, medical history, medications, symptoms, and laboratory results.
The proposed framework envisions an AI-driven system that would provide clear, written outputs useful for both healthcare providers and patients. By holistically evaluating all elements of a patient’s situation, such a system could potentially deliver more personalized diagnoses and treatments while supporting clinician decision-making rather than replacing it.
It is important to emphasize that 3RDI is designed as a clinical decision support tool and not a medical device. As such, it operates within the regulatory framework that distinguishes clinical decision support software from medical devices, allowing for independent review and final decision-making by healthcare professionals. All recommendations generated by the system would require review and approval by qualified clinicians, maintaining the healthcare provider as the ultimate decision-maker in the diagnostic and treatment process.
Conceptual Framework Design
The 3RDI framework envisions a system architecture designed to balance advanced AI capabilities with practical clinical integration. This section outlines the key components of this conceptual framework, including data handling, processing approaches, output structuring, and feedback mechanisms.
Input Data Architecture and Target Population
The proposed 3RDI framework would process comprehensive patient data structured in standardized formats such as XML (eXtensible Markup Language) or JSON (JavaScript Object Notation). To capture a complete clinical picture, the system would incorporate categories of information as listed in Table 3.
These elements were selected based on input from medical professionals experienced in UTI diagnosis and treatment. Appendix A and Appendix B illustrate examples of how medical history and daily logs might be structured in an XML format for processing by the system.
The framework also envisions integration capabilities with electronic medical record (EMR) systems such as Epic, allowing patient data to be retrieved directly from existing clinical systems rather than requiring manual entry.
Target Population
The 3RDI framework is designed to support specific patient and healthcare provider populations:
Patients
Adult patients presenting with symptoms suggestive of UTI in inpatient and outpatient settings are the targets of the system. The framework would be particularly valuable for complex cases involving comorbidities, recurrent infections, or atypical presentations where diagnostic uncertainty is higher. While the initial conceptual framework focuses on adult patients, future iterations could be adapted for pediatric populations with appropriate modifications to account for different clinical presentations, diagnostic criteria, and treatment considerations in children.
Healthcare providers, the primary users of the 3RDI system, would be healthcare providers involved in UTI diagnosis and management, including primary care physicians, emergency department clinicians, hospitalists, infectious disease specialists, and urologists. The framework is designed to be most valuable for providers with high patient volumes or those practicing in settings without immediate access to specialist consultation, where decision support could help standardize care according to best practices.
Proposed AI Processing Approach
The 3RDI conceptual framework proposes using a specialized LLM that is trained and fine-tuned for medical applications, particularly UTI diagnosis and treatment. This approach offers several theoretical advantages (Table 4).
For initial development of this framework, synthetic UTI patient data would be used rather than real patient records. This approach minimizes ethical concerns related to patient privacy while allowing for rapid prototyping and iterative improvements. The framework envisions a development process where synthetic training data would be curated in collaboration with domain experts—including physicians and medical professionals—to ensure clinical relevance and accuracy.
The prototype development would include creating structured synthetic patient profiles based on common clinical presentations, symptoms, and treatment pathways for UTIs. A panel of experienced physicians would review and refine these examples to ensure alignment with real-world medical practices, resulting in a dataset that closely mirrors real-world clinical scenarios.
Patient data would be stored in a standardized format, allowing the model to recognize patterns in patient symptoms, laboratory results, and other medical history details, ultimately generating reliable diagnostic and treatment suggestions. After initial training, the model would be fine-tuned using validated medical literature on UTIs, treatment protocols, and clinical guidelines.
The DETNQ Output Framework
A core innovation in the 3RDI conceptual framework is the DETNQ structure for organizing clinical outputs. This framework is designed to transform complex medical data into actionable clinical decisions while addressing key limitations in traditional CDSS outputs. Table 5 compares the DETNQ framework with other common CDSS output structures.
Diagnosis
This segment would present primary and differential diagnoses in order of likelihood, clearly delineating between confirmed and suspected conditions. It would integrate relevant International Classification of Diseases, 10th Revision (ICD-10) codes for accurate documentation and highlight critical findings requiring immediate attention.
Evidence
The Evidence component would offer a hierarchical presentation of supporting clinical data, linking symptoms, laboratory results, and diagnostic conclusions. It would identify pattern matches with historical cases, conduct quantitative analyses of diagnostic confidence based on available data, and integrate relevant clinical guidelines and best practices. Figure 1 shows a conceptual example of the Evidence component.
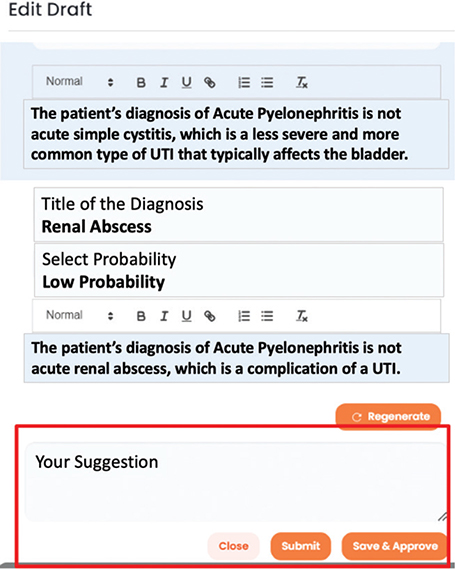
Fig. 1. A conceptual example of the Evidence component in the DETNQ output framework. DETNQ: Diagnosis, Evidence, Treatment Plan, Notes, Quality.
Treatment Plan
This section would provide prioritized intervention recommendations, including medication suggestions with dosing considerations and alternative treatment options based on patient-specific factors. It would issue contraindication warnings, drug interaction alerts, and timelines for expected clinical improvement while setting criteria for treatment modification or escalation. Appendix C illustrates a proposed Treatment Plan output.
Notes
The Notes section would include context-specific clinical pearls and considerations, documentation guidance for billing compliance, relevant research findings and clinical trial data, patient education materials, discharge instructions, follow-up recommendations, and monitoring parameters. Appendix D shows an example of how the Notes section might appear.
Quality
Finally, the Quality segment would ensure alignment with relevant quality measures, such as Healthcare Effectiveness Data and Information Set (HEDIS) and Merit-based Incentive Payment System (MIPS), assess documentation completeness, monitor antibiotic stewardship compliance metrics, calculate risk stratification scores, and provide care coordination recommendations. Appendix E provides a conceptual example of the Quality component output.
This daily iterative process would be designed to align with the clinical management of patients, reflecting the workflow of daily clinical rounds. By mimicking clinical patient evaluation and presenting information in a familiar structure, the framework aims to increase potential clinician acceptance. The DETNQ approach could help address workflow disruption and lack of clinical engagement, which are primary deterrents to clinician adoption of CDSS.21
The DETNQ framework would be enhanced by a confidence scoring system, which would assign numerical metrics to each recommendation. This scoring system would operate on a scale of 0–100, with 100 representing the highest confidence recommendations and 0 representing the lowest. The confidence score would provide additional context to clinicians as they evaluate recommendations, helping them determine when additional review or alternative approaches might be warranted.
Continuous Learning and Clinician Feedback
A fundamental innovation of the 3RDI framework is its proposed approach to continuous learning through clinician feedback. Unlike static CDSS that remain unchanged after deployment, 3RDI would incorporate a structured feedback loop to ensure ongoing improvement and adaptation to clinical realities. The envisioned feedback mechanism is presented in Table 6.
Figure 1 illustrates a conceptual user interface for this feedback mechanism, showing how clinicians might edit the model’s output to improve accuracy.
This approach would not only allow subject-matter experts to correct inaccurate outputs but also provide an opportunity to personalize the model to the specifications of different user groups or institutions. For example, some hospitals or clinicians might prefer to prescribe cefepime instead of ceftazidime for patients with UTIs. The 3RDI model would be designed to recognize such patterns over time and make adjustments for specific user groups.
The complete user journey, from initial login to receiving AI-assisted recommendations, is conceptualized in Figure 2, which visualizes a nine-step process from the platform’s login page to receiving the model response in DETNQ format for a new patient.
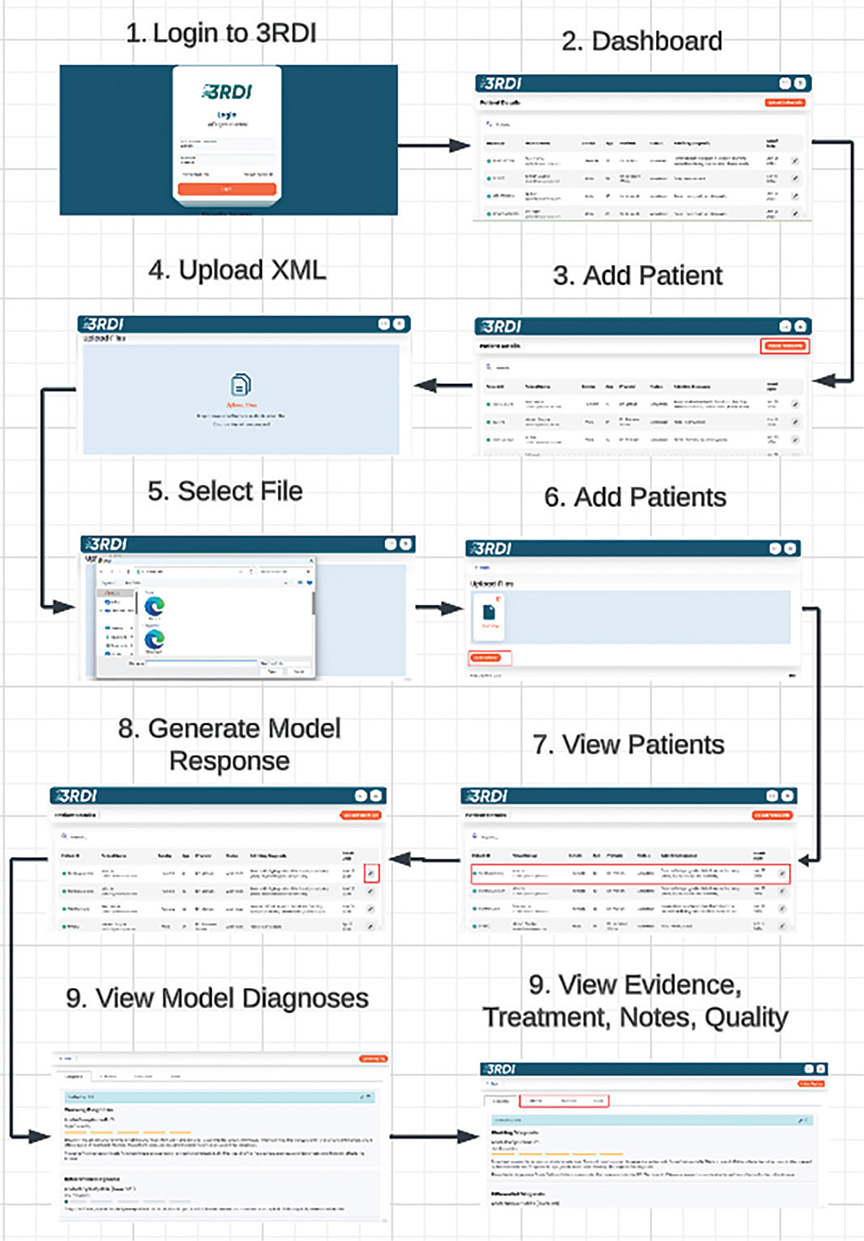
Fig. 2. A nine-step process from the platform’s login page to receiving the model response in DETNQ format for a new patient. 3RDI: an automatic photographic apparatus; DETNQ: Diagnosis, Evidence, Treatment Plan, Notes, Quality; XML: eXtensible Markup Language.
Potential Innovations, Benefits, and Limitations
The 3RDI conceptual framework introduces several potential innovations that could address existing challenges in CDSS, particularly for UTI diagnosis and treatment. This section explores the theoretical benefits and practical limitations of implementing such a framework in real-world clinical settings.
Day-wise Iterative Approach
This approach would align with the natural workflow of daily patient rounds, potentially allowing dynamic treatment adjustments based on patient responses while reducing cognitive load by presenting information in familiar clinical patterns.
Real-time Clinical Adaptation
Such a system could potentially adjust automatically to institutional antibiotic preferences, learn from local resistance patterns, and tailor recommendations for specific patient populations. By adapting to resource availability, facility capabilities, and regional healthcare guidelines, the framework could be applicable across diverse clinical settings.
Continuous Learning System
As a CLS, 3RDI would incorporate real-time model updates based on clinical feedback, making it potentially adaptable to evolving clinical knowledge and standards. The CLSs in healthcare enable models to adapt and improve, potentially enhancing diagnostic precision.22 The ability to automatically incorporate new medical evidence and dynamically adjust to changing clinical patterns could help ensure that the diagnostic process remains accurate and reliable.
Human-AI Collaboration
Integrating human expertise with AI-driven systems offers a potentially powerful synergy that could maximize accuracy in diagnosis and decision-making. The use of AI-driven CDSS in conjunction with human expertise has been shown to reduce laboratory workloads and decrease the need for unnecessary urine cultures and antibiotic prescriptions.23 While human clinicians bring contextual understanding, intuition, and experience, AI systems excel in processing vast amounts of data and identifying patterns that may be difficult for humans to detect.
Clinical Confidence Metrics
The inclusion of confidence metrics, such as the proposed perplexity score system, could provide transparency in AI decision-making, potentially aiding clinicians in prioritizing cases for review, stratifying risk in complex scenarios, and determining when to escalate care.
Potential Healthcare System Benefits
If successfully implemented, the 3RDI framework could potentially offer several system-wide improvements:
Workflow Optimization
A properly designed system might reduce time spent on UTI diagnosis and documentation, streamline clinical decision-making, improve care team communication, reduce administrative burdens, and enhance clinical documentation efficiency.
Resource Allocation Improvements
Theoretical benefits could include a reduction in laboratory testing, optimization of antibiotic usage patterns, better utilization of specialist consultations, fewer emergency department visits, and more efficient use of nursing resources.
Cost Reduction through Accuracy
Potential economic benefits might include decreased preventable UTI-related hospitalizations, shorter lengths of stay for admitted patients, reduced readmission rates, lower costs related to antibiotic resistance, and improved reimbursement driven by better documentation.
Quality and Safety Enhancements
Such a system could potentially reduce antibiotic prescription errors, improve compliance with clinical guidelines, enhance adherence to quality measures, increase patient satisfaction, and reduce medical-legal risk through standardized documentation.
Implementation Challenges
Practical implementation of the 3RDI framework would face several significant challenges:
Technical Integration
Integration with existing electronic health record (EHR) systems remains complex, requiring careful API development, data mapping, and workflow analysis. Different healthcare institutions use various EHR platforms with different data structures, necessitating flexible integration approaches.
Computational Requirements
While cloud-based deployment options could mitigate hardware limitations, ensuring reliable, low-latency performance across diverse healthcare settings would require careful architecture planning and resource allocation.
Training Data Diversity
Ensuring the system performs equitably across diverse patient populations would require careful attention to training data composition. Biases in training data could lead to disparate performance across demographic groups.
Regulatory Navigation
AI-driven clinical decision support tools face evolving regulatory requirements. The framework would need to be implemented with clear pathways for regulatory compliance, including appropriate validation studies and documentation.
Change Management
Perhaps the most challenging would be managing the cultural change required for clinician adoption. Resistance to AI-assisted tools remains common, requiring thoughtful implementation strategies that emphasize augmentation rather than replacement of clinical judgment.
Limitations of the Proposed Framework
Several important limitations must be acknowledged in the 3RDI conceptual framework:
AI Hallucination and Uncertainty
Like all LLMs, any implementation of this framework could potentially generate responses based on incomplete or ambiguous input data. To mitigate this, confidence scoring and clinician oversight mechanisms would be essential components of any implementation.
Need for Clinical Validation
The framework remains theoretical and would require extensive clinical validation before deployment. Future implementations would need prospective studies and randomized controlled trials to ensure the robustness and safety of any recommendations.
Adoption and Provider Trust Challenges
Clinicians may hesitate to rely on AI-generated recommendations, fearing misinterpretations or biases in decision-making. Any implementation would need to incorporate clinician-in-the-loop learning, allowing providers to modify outputs and improve model accuracy over time.
Regulatory Status
The 3RDI is conceptualized as a clinical decision tool rather than a medical device, meaning it serves to augment clinical judgment rather than replace it. All recommendations would require review and approval by qualified healthcare professionals, maintaining the clinician as the ultimate decision-maker in the diagnostic and treatment process. This design approach aligns with current regulatory distinctions between CDSS and medical devices.
Privacy and Security Requirements
Handling sensitive patient data would require robust security measures and compliance with regulations such as HIPAA. Any implementation would need careful design of data protection mechanisms, access controls, and audit trails.
Bias Mitigation and Health Equity Considerations
The potential for AI systems to perpetuate or amplify existing healthcare disparities represents a significant ethical concern that must be proactively addressed in any implementation of the 3RDI framework. While our conceptual design includes clinician feedback as one mechanism to correct model outputs, we recognize this approach alone is insufficient to ensure equitable performance across diverse patient populations.
A comprehensive approach to bias mitigation would require multiple strategies implemented throughout the development and deployment process (Table 7).
The framework would also include a “fairness dashboard” that continuously monitors system performance across different patient populations, flagging any emerging disparities for immediate attention. This multilayered approach acknowledges that addressing health equity in AI systems requires intentional design choices at every stage of development and implementation, rather than relying solely on post-hoc corrections by clinicians.
Future Research Directions and Technical Implementation Considerations
The 3RDI conceptual framework presents several avenues for future research and development. Future work should include rigorous validation using both retrospective and prospective approaches. Initially, retrospective validation using anonymized real-world patient data would provide insights into the framework’s potential performance. This would be followed by prospective clinical trials comparing AI-assisted diagnosis and treatment with standard care approaches. What a proposed technical implementation would involve is shown in Appendix F.
This pseudocode represents a high-level implementation approach that could be further developed into a working prototype for validation studies. The actual implementation would require careful design of each component, particularly:
Prompt Engineering Approach
Methods for constructing effective medical prompts that elicit structured reasoning from the LLM while maintaining clinical relevance. This includes designing prompts that encourage the model to follow the DETNQ structure while incorporating relevant medical knowledge.
Context Window Management
Techniques for efficiently processing comprehensive patient histories within the limited context windows of current LLMs. This includes prioritization algorithms for selecting the most relevant patient information when dealing with complex cases.
Confidence Score Calculation
Mathematical approaches to quantifying confidence based on model perplexity, reasoning consistency, and evidence strength. This would include developing metrics that correlate with clinical accuracy.
Feedback Integration Mechanisms
Technical approaches for processing clinician feedback and incorporating it into model improvement, both for immediate response refinement and long-term learning.
Framework Extension
While initially focused on UTIs, the underlying architecture could potentially be extended to other infectious diseases and clinical conditions. Research into the transferability of the DETNQ structure and feedback mechanisms to other domains would be valuable. Figure 2 shows the proposed system architecture with data flows between components, providing a visual representation of how the various elements of the framework would interact in a potential implementation.
Validation Methodology Development
Establishing standardized methodologies for evaluating AI-driven CDSS remains an ongoing challenge. Future research should focus on developing specific validation protocols that assess both technical performance metrics (such as diagnostic accuracy and treatment appropriateness) and clinical utility measures (such as workflow integration, user satisfaction, and impact on patient outcomes).
Such protocols might include (1) comparative performance testing (multiarmed trials comparing AI-assisted diagnosis with traditional clinical pathways and other decision support tools), (2) simulation studies (using synthetic patient cases with known ground truth to assess diagnostic accuracy across diverse presentations), (3) user experience evaluation (structured assessments of clinician interaction patterns, cognitive load, and satisfaction with system outputs), and (4) long-term impact assessment (longitudinal studies evaluating changes in clinical practice patterns, antibiotic stewardship metrics, and patient outcomes following system implementation).
Model Explainability Enhancements
Developing improved methods for making AI reasoning transparent and understandable to clinicians represents an important research direction. This could include visual explanations of diagnostic reasoning and evidence weighting. Future work might explore:
Future work might explore (1) Evidence visualization tools (graphical representations of the relationship between symptoms, laboratory values, and diagnostic conclusions), (2) confidence attribution (methods for communicating which specific data elements most influenced the system’s recommendations), (3) alternative scenario analysis (tools that allow clinicians to explore how different clinical findings might impact recommendations), and finally, (4) diagnostic reasoning pathways (visual representation of the logical steps from data to conclusion).
Bias Mitigation Strategies
Research into specific techniques for identifying and mitigating biases in clinical AI systems would strengthen implementations of this framework. This includes both technical approaches to bias detection and process improvements in model development and validation.
Priority research areas include Demographic Performance Auditing (methods for systematically assessing system performance across different patient populations), Fairness-Aware Training Techniques (algorithmic approaches that balance accuracy with equitable performance across groups), Synthetic Data Generation (techniques for augmenting training data to improve representation of underrepresented groups, and Continuous Monitoring Frameworks (methods for detecting emergent biases as system performance evolves).
Conclusion
The 3RDI conceptual framework represents a forward-looking approach to AI-driven CDSS for conditions such as UTIs. By proposing the integration of LLMs, ML algorithms, and iterative feedback loops, this framework envisions systems that could enhance the diagnostic process, offering personalized, real-time recommendations that align with each patient’s unique medical profile.
The day-wise iterative approach proposed in the framework mirrors the natural workflow of clinical rounds, potentially enabling dynamic updates to diagnoses and treatment plans as new patient data becomes available. This temporal processing approach represents a significant conceptual advance beyond traditional static CDSS models, suggesting a pathway to continuous, adaptive response to the evolving nature of patient care.
A key innovation lies in the framework’s CLS concept, which incorporates real-time clinician feedback to refine algorithms. Unlike traditional CDSS requiring periodic manual updates, this feedback loop could facilitate immediate model improvement and institution-specific customization, addressing the limitations of standardized, nonadaptable recommendations. This collaborative dynamic between AI and clinicians could potentially bolster clinical accuracy while enhancing efficiency in decision-making.
Additionally, the DETNQ structure introduced by the framework provides an organized output format that would support clinical decision-making while ensuring compliance with quality measures. The inclusion of confidence scoring, quality metrics, and standardized documentation guidance represents a conceptual advancement over existing CDSS outputs, addressing critical needs for transparency, confidence assessment, and regulatory alignment.
The potential benefits of implementing the 3RDI framework extend beyond UTI management, with possibilities for streamlining clinical workflows by reducing unnecessary laboratory tests and antibiotic prescriptions, thereby supporting antibiotic stewardship. If successfully implemented and validated, such systems could be extended to other infectious diseases, establishing a scalable approach for comprehensive AI-driven CDSS.
By advancing a vision for precision medicine through the evolving synergy of AI and human expertise, the 3RDI framework proposes a direction for intelligent, responsive healthcare support systems that could optimize resources while improving patient outcomes. However, significant research, validation, and implementation work remains to be done before these theoretical benefits can be realized in clinical practice.
Our approach explicitly acknowledges the importance of addressing health equity considerations and bias mitigation through comprehensive strategies that go beyond simply relying on clinician feedback. By incorporating rigorous technical and governance approaches to ensure fair performance across diverse populations, the framework seeks to advance health equity alongside clinical efficiency and accuracy.
As we look into the future, the 3RDI framework offers a blueprint for how AI and human expertise might work together to enhance healthcare delivery for one of medicine’s most common conditions. The concepts presented here may help guide the development of next-generation CDSS that are not only more intelligent but also more adaptable, transparent, and equitable than current approaches.
Guidelines might include that the research involves no more than minimal risk to the subjects, could not practicably be carried out without the requested waiver or alteration, and involves using identifiable private information or identifiable biospecimens, the research could not practicably be carried out without using such information or biospecimens in an identifiable format. In addition, the waiver or alteration will not adversely affect the rights and welfare of the subjects, and whenever appropriate, the subjects or legally authorized representatives will be provided with additional pertinent information after participation.
Conflicts of Interest
Author Dr. Amar Gupta is an Editorial Board Member of THMT. He has disqualified himself from the peer review and decision-making process.
Contributors
Manoj Jain: Conceptualization, methodology, writing, review, editing, supervision, project administration. Hiren Pokharna: Methodology, data curation. Sridhar Sunkara: Software, validation, formal analysis, data curation. Sanjeev Bora: Software, validation, formal analysis, visualization, review, editing. Kiran Ponamgi: Software, validation, data curation, visualization. Rohan Dang Sharma: Investigation, resources, writing, review, editing. Amar Gupta: Conceptualization, supervision, project administration, review & editing. All authors have read and agreed to the published version of the manuscript.
Data Availability Statement (DAS), Data Sharing, Reproducibility, and Data Repositories
The data that support the findings of this study are available from the corresponding author, Manoj Jain, upon reasonable request.
Application of AI-Generated Text or Related Technology
AI was used to assist in creating the abstract and plain language summary.
Acknowledgments
None
References
- Mebrahtu TF, Skyrme S, Randell R, Keenan A-M, Bloor K, Yang H, et al. Effects of computerised clinical decision support systems (CDSS) on nursing and allied health professional performance and patient outcomes: a systematic review of experimental and observational studies. BMJ Open. 2021;11(12):e053886. https://doi.org/10.1136/bmjopen-2021-053886
- Zhao C, Liang N, Zhang H, Li H, Yang Y, Zong X, et al. Harnessing the power of clinical decision support systems: challenges and opportunities. Open Heart. 2023;10(2):e002432. https://doi.org/10.1136/openhrt-2023-002432
- Ozkan IA, Koklu M, Sert IU. Diagnosis of urinary tract infection based on artificial intelligence methods. Comput Methods Programs Biomed. 2018;166:51–59. https://doi.org/10.1016/j.cmpb.2018.10.007
- Elhaddad M, Hamam S. AI-driven clinical decision support systems: an ongoing pursuit of potential. Cureus. 2024;16(4):e57728. https://doi.org/10.7759/cureus.57728
- Naik N, Talyshinskii A, Shetty DK, Hameed BMZ, Zhankina R, Somani BK. Smart diagnosis of urinary tract infections: is artificial intelligence the fast-lane solution? Curr Urol Rep. 2023;25(1):37–47. https://doi.org/10.1007/s11934-023-01192-3
- Xu R, Deebel N, Casals R, Dutta R, Mirzazadeh M. A new gold rush: a review of current and developing diagnostic tools for urinary tract infections. Diagnostics. 2021;11(3):479. https://doi.org/10.3390/diagnostics11030479
- Hojat LS, Saade EA, Hernandez AV, Donskey CJ, Deshpande A. Can electronic clinical decision support systems improve the diagnosis of urinary tract infections? A systematic review and meta-analysis. Open Forum Infect Dis. 2022;10(1):ofac691. https://doi.org/10.1093/ofid/ofac691
- McDermott KW, Jiang HJ. Characteristics and costs of potentially preventable inpatient stays, 2017. Rockville, MD: Agency for Healthcare Research and Quality (US); 2006.
- Burton RJ, Albur M, Eberl M, Cuff SM. Using artificial intelligence to reduce diagnostic workload without compromising detection of urinary tract infections. BMC Med Inform Decis Mak. 2019;19(1):171. https://doi.org/10.1186/s12911-019-0878-9
- Computer-Based Medical Consultations: MYCIN. In: Edward H. Shortliffe editors. New York: Elsevier; 1976. 286 pp. ISBN-10: 0444569693. ISBN-13: 978-0444569691
- Berner ES. Clinical decision support systems: theory and practice. Springer Science & Business Media; (2nd ed.). NewYork: Springer Verlag; 2007.
- de Vries S, Doesschate TT, Totté JEE, Heutz JW, Loeffen YGT, Oosterheert JJ, et al. A semi-supervised decision support system to facilitate antibiotic stewardship for urinary tract infections. Comp Biol Med. 2022;146:105621. https://doi.org/10.1016/j.compbiomed.2022.105621
- Gupta GK, Singh A, Manikandan SV, Ehtesham A. Digital diagnostics: the potential of large language models in recognizing symptoms of common illnesses. AI. 2025;6(1):13. https://doi.org/10.3390/ai6010013
- Ong JCL, Jin L, Elangovan K, Lim GYS, Lim DYZ, Sng GGR, et al. Development and testing of a novel large language model-based clinical decision support systems for medication safety in 12 clinical specialties. arXiv preprint arXiv:2402.01741, 2024 arxiv.org.
- Liu S, Wright AP, Patterson BL, Wanderer JP, Turer RW, Nelson SD, et al. Using AI-generated suggestions from ChatGPT to optimize clinical decision support. J Am Med Inform Assoc. 2023;30(7):1237–45. https://doi.org/10.1093/jamia/ocad072
- Silcox C, Dentzer S, Bates DW. AI-enabled clinical decision support software: a “trust and value checklist” for clinicians. NEJM Catalyst. 2020;1(6). https://doi.org/10.1056/cat.20.0212
- Sutton RT, Pincock D, Baumgart DC, Sadowski DC, Fedorak RN, Kroeker KI. An overview of clinical decision support systems: benefits, risks, and strategies for success. NPJ Digit Med. 2020;3:17. https://doi.org/10.1038/s41746-020-0221-y
- Richardson C, Robb KA, O’Connor RC. A systematic review of suicidal behaviour in men: A narrative synthesis of risk factors. Soc Sci Med. 2021;276:113831. https://doi.org/10.1016/j.socscimed.2021.1138
- Wang L, Zhang Z, Wang D, Cao W, Zhou X, Zhang P, et al. Human-centered design and evaluation of AI-empowered clinical decision support systems: a systematic review. Front Comput Sci. 2023;5:1187299. https://doi.org/10.3389/fcomp.2023.1187299
- Gomez-Cabello CA, Borna S, Pressman S, Haider SA, Haider CR, Forte AJ. Artificial-intelligence-based clinical decision support systems in primary care: a scoping review of current clinical implementations. Eur J Investig Health Psychol Educ. 2024;14(3):685–98. https://doi.org/10.3390/ejihpe14030045
- Laka M, Milazzo A, Merlin T. Factors that impact the adoption of clinical decision support systems (CDSS) for antibiotic management. Int J Environ Res Public Health. 2021;18(4):1901. https://doi.org/10.3390/ijerph18041901
- Flores E, Martínez-Racaj L, Blasco Á, Diaz E, Esteban P, López-Garrigós M, et al. A step forward in the diagnosis of urinary tract infections: from machine learning to clinical practice. Comput Struct Biotechnol J. 2024;24:533–41. https://doi.org/10.1016/j.csbj.2024.07.018
- Sahiner B, Friedman B, Linville C, Ipach C, Montgomery E, Alexander ES, et al. Perspectives and best practices for artificial intelligence and continuously learning systems in healthcare. Cincinnati, OH: Xavier University Library; 2018.
- Talyshinskii A, Naik N, Hameed BMZ, Juliebø-Jones P, Somani BK. Potential of AI-driven chatbots in urology: revolutionizing patient care through artificial intelligence. Curr Urol Rep. 2024;25(1):9–18. https://doi.org/10.1007/s11934-023-01184-3
- Labkoff S, Oladimeji B, Kannry J, Solomonides A, Leftwich R, Koski E, et al. Toward a responsible future: recommendations for AI-enabled clinical decision support. J Am Med Inform Assoc. 2024;31(11):2730–9. https://doi.org/10.1093/jamia/ocae209
Copyright Ownership: This is an open-access article distributed in accordance with the Creative Commons Attribution Non-Commercial (CC BY-NC 4.0) license, which permits others to distribute, adapt, enhance this work non-commercially, and license their derivative works on different terms, provided the original work is properly cited and the use is non-commercial. See http://creativecommons.org/licenses/by-nc/4.0. The authors of this article own the copyright.
Appendix
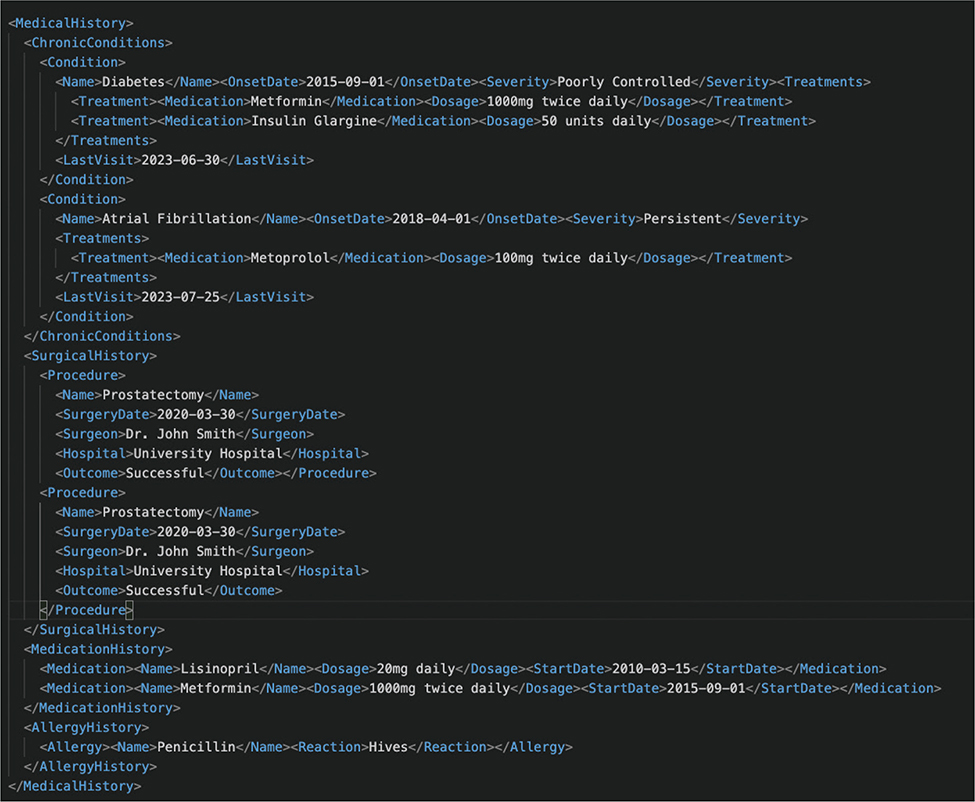
Appendix A. An example of medical history XML (eXtensible Markup Language).
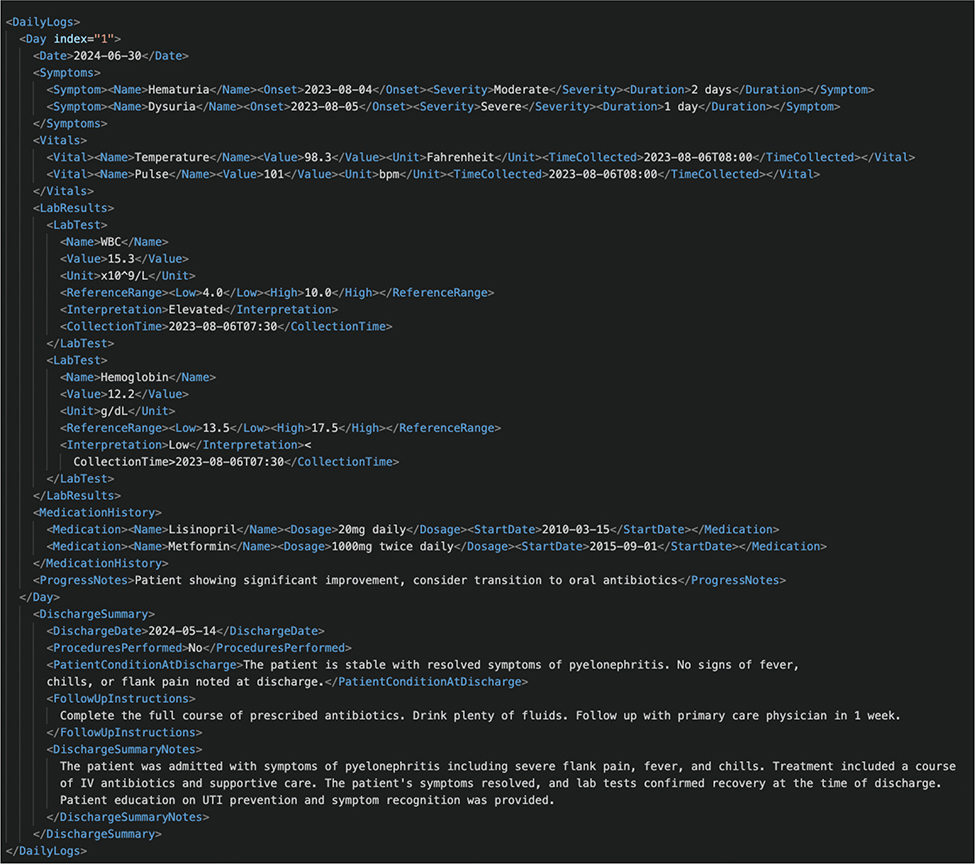
Appendix B. An example of daily logs XML (eXtensible Markup Language).
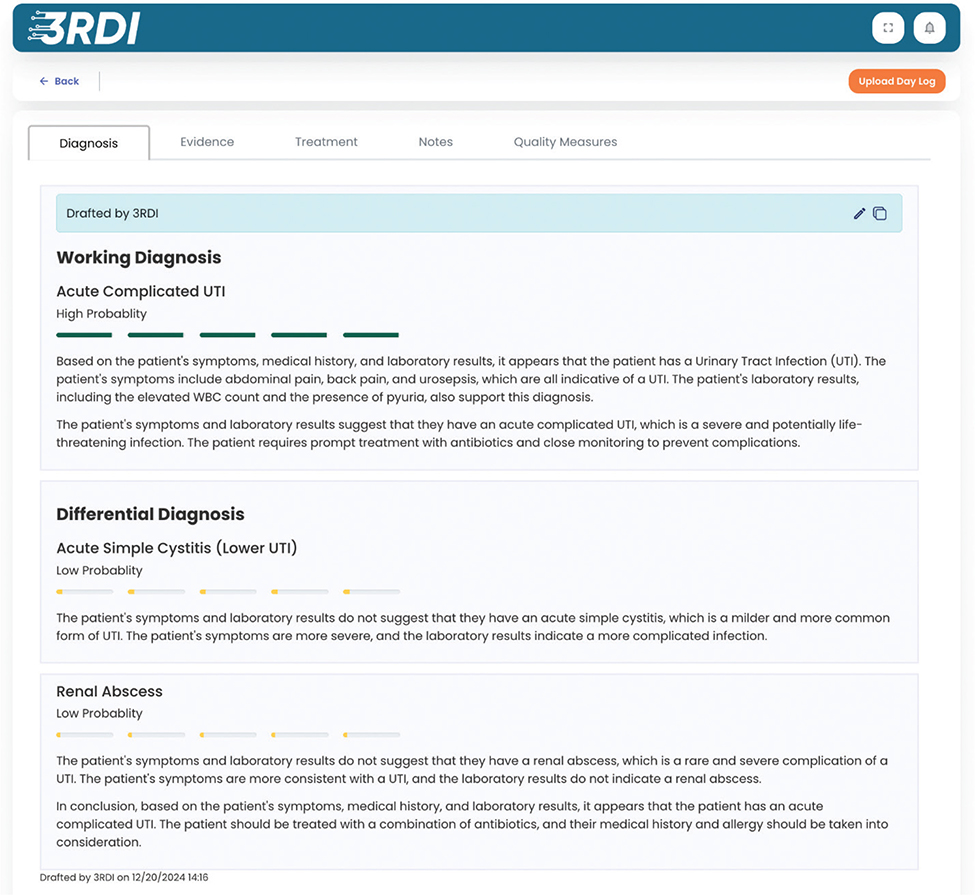
Appendix C. An example of how the Diagnosis component might appear in the proposed system.
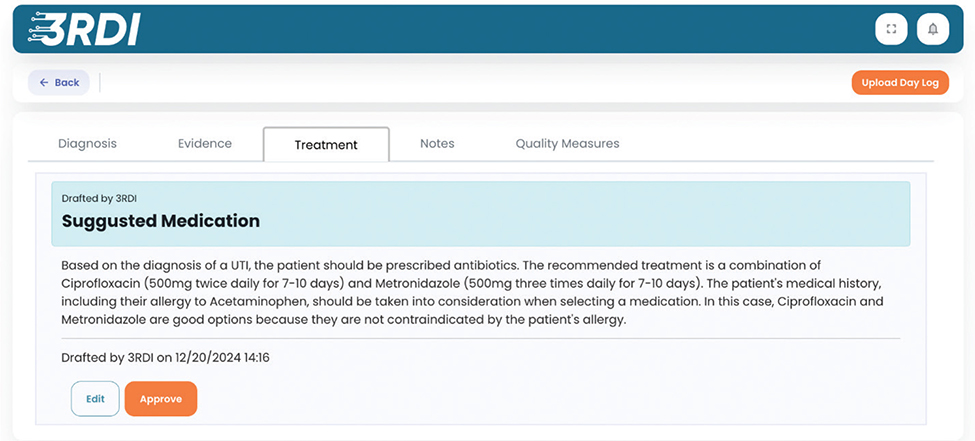
Appendix D. A proposed Treatment Plan output.
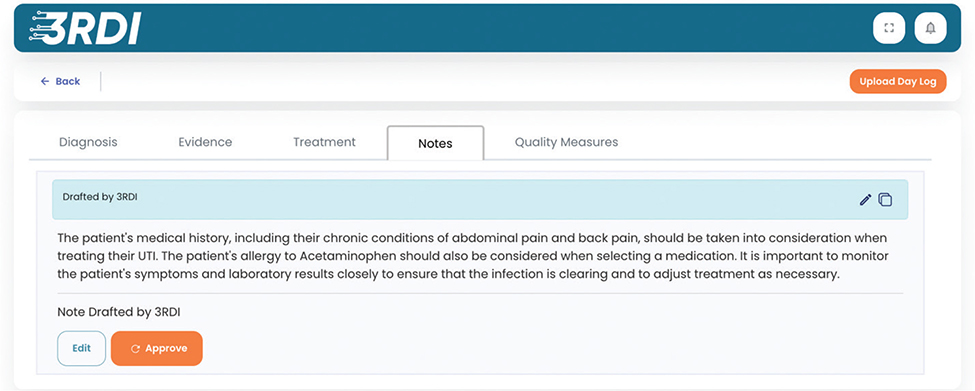
Appendix E. An example of how the Notes section might appear.
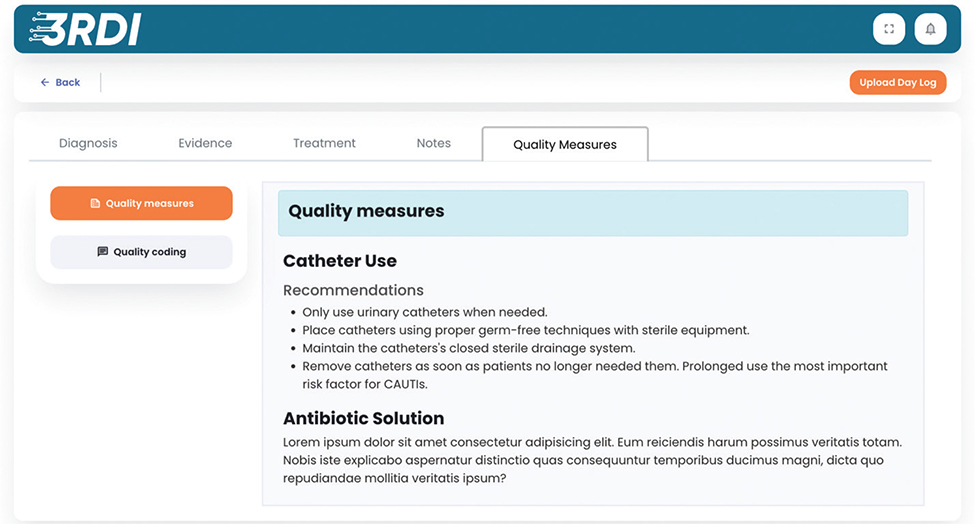
Appendix F. A conceptual example of the Quality component output.