ORIGINAL RESEARCH
An Explainable Deep Transfer Learning Approach with Augmentation for Chest X-Ray-Driven Pulmonary Disease Diagnosis
R. Sriramkumar, M.Tech1  , K. Selvakumar, PhD1
, K. Selvakumar, PhD1  and J. Jegan, PhD2
and J. Jegan, PhD2 
1Department of Information Technology, Annamalai University, Chidambaram, India; 2Department of Computer Science and Engineering, SRM Institute of Science and Technology, Tiruchirappalli, Tamil Nadu, India
Keywords: Chest X-ray, clinical decision support, ConvNeXt, deep transfer learning, EfficientNetV2, ensemble learning, pulmonary disease detection, Swin Transformer
Abstract
Objective: Chest X-ray (CXR) images are a fundamental diagnostic tool for pulmonary diseases; however, their interpretation is often hindered by observer variability, scarce annotated sets, and the requirement for timely clinical judgment.
Methods: To address these issues, the authors applied an augmented deep transfer learning framework that synthesizes three state-of-the-art architectures, namely, EfficientNetV2, Convolutional Neural Networks for Image Classification (ConvNeXt), and Swin Transformer, into a hybrid ensemble to enhance diagnostic consistency and precision. Each model has distinct strengths: EfficientNetV2 provides a lightweight, scalable representation. ConvNeXt contributes an up-to-date convolutional structure, and Swin Transformer extracts long-range relationships using hierarchical attention mechanisms. For augmentation, MixUp, CutMix, and RandAugment were used to augment robustness, and stratified sampling and focal loss were used to reduce class imbalance. An annotated COVID-CXR dataset of three classes—COVID-19, pneumonia, tuberculosis, lung cancer, pleural effusion, and normal—was used to examine the framework.
Results: The ensemble model outperformed the individual baseline models, achieving an overall precision of 99.3%, sensitivity of 99.0%, and specificity of 99.2%, with an area under the curve above 0.98. Visualization by gradients validated that the system systematically emphasized clinically significant lung areas, supporting its interpretability for medical application.
Conclusion: The results substantiate that combining advanced convolutional and transformer-based architecture designs, bolstered by augmentation and interpretability, can significantly enhance CXR diagnostics. Future research will aim to extend to larger multicenter sets, merge complementary imaging modalities, and devise deployment methodologies appropriate for varied healthcare settings.
Plain Language Summary
Chest X-rays (CXR) are commonly used to detect lung diseases, but manual interpretation can vary. This study presents a deep learning model combining EfficientNetV2, ConvNeXt, and Swin Transformer to identify diseases such as COVID-19, pneumonia, tuberculosis, and others from CXR. Data augmentation (MixUp, CutMix, and RandAugment) and focal loss improved accuracy and fairness across disease types. The system achieved 99.3% accuracy and visualized affected lung areas for interpretability, supporting clinical trust. The model can help radiologists diagnose faster and improve healthcare access, especially in low-resource hospitals and telemedicine applications.
Citation: Telehealth and Medicine Today © 2025, 10: 594
DOI: https://doi.org/10.30953/thmt.v10.594
Copyright: © 2025 The Authors. This is an open-access article distributed in accordance with the Creative Commons Attribution Non-Commercial (CC BY-NC 4.0) license, which permits others to distribute, adapt, and enhance this work non-commercially, and license their derivative works on different terms, provided the original work is properly cited and the use is non-commercial. See http://creativecommons.org/licenses/by-nc/4.0. The authors of this article own the copyright.
Received: May 27, 2024; Accepted: October 14, 2025; Published: December 17, 2025
Corresponding Author: R. Sriramkumar Mail Id: sriramkumar2686@gmail.com
Competing interests and funding: No competing interests exist among the authors. The authors declare that they have no financial or non-financial relationships or activities that could have influenced the results or interpretation of this research.
This research did not receive any specific grant from funding agencies in the public, commercial, or not-for-profit sectors. This work was carried out as part of the author’s doctoral research under Annamalai University.
Chest radiography, by virtue of its universal availability, economy, and noninvasive nature, has historically been the underlying foundation of thoracic diagnostics, serving as the first strategy in the identification of pulmonary diseases that extend from acute infections to chronic malignancies. Despite its importance, the interpretation process itself is marked by significant inter-observer variability, inherent subjectivity, and diagnostic disagreements that often delay the initiation of treatment protocols. These pitfalls are even more pronounced in situations characterized by large patient volumes, restricted resources, or limited access to expert radiologic knowledge.
Over the past few years, the growth of data-based computational frameworks has markedly changed the landscape of medical image analysis by offering methodologies capable of unveiling latent hierarchical features that transcend the perceptual abilities of the human visual cortex. Among these approaches, deep transfer learning models, which fine-tune pre-trained models that were originally trained over large generic datasets for specific domain applications, are most relevant for addressing the challenges of sparse annotated medical datasets, particularly for chest radiography, where large and representative datasets with different pathologies (COVID-19, viral and bacterial pneumonia, tuberculosis (TB), pulmonary tumors, pleural effusions, and interstitial abnormalities) are rarely available in large numbers.
At the same time, extending convolutional architectures to new models (e.g. convolutional neural networks for image classification (ConvNeXt) and EfficientNetV2) and combining them with hierarchical attention mechanisms (notably the Swin Transformer) have opened new paths for image classification. These networks take advantage of the fine-grained nature of local convolutional filters and the ability to model long-range contextual relationships, both of which are particularly beneficial for the varied and often subtle manifestations of thoracic disease. Combined with state-of-the-art preprocessing techniques such as contrast enhancement, adaptive normalization, and aggressive augmentation techniques such as MixUp, CutMix, and RandAugment, these networks are highly robust to class imbalance, overfitting, and domain shifts.
In addition to numerical dominance, the clinical translatability of such systems depends on their interpretability, ethical acceptability, and assimilation into mainstream diagnostic workflows. Gradient-based saliency maps and class-discriminative visualization can serve as an adjunct to provide clinicians with a means of verifying that the computational region of interest is consistent with radiologically salient regions, thereby building trust and responsible computer-assisted decision-making.
Against this background, these researchers developed an enhanced deep transfer learning model that employs a hybrid ensemble of EfficientNetV2, ConvNeXt, and Swin Transformer to address the problem of pulmonary disease detection from chest radiographs. The goal of this research is (1) to demonstrate enhanced diagnostic performance across a wide range of disease categories, (2) to achieve robustness using augmentation and imbalance management techniques, and (3) to produce interpretable and clinically relevant results. This development holds the promise of computationally enhanced chest radiography as a valuable adjunct to standard diagnostic practice, especially in low-resource, high-need clinical settings.
Literature Survey
Appendix A lists the contributions by researchers since 2022 to develop an enhanced deep transfer learning model that employs a hybrid ensemble of EfficientNetV2, ConvNeXt, and Swin Transformer to address the problem of pulmonary disease detection from chest radiographs.
The integration of artificial intelligence (AI) into real-world workflows requires careful consideration of safety, ethics, and clinical feasibility. In a study that explored the application of image-based computational systems in surgical practice, Nguyen et al.14 identified both efficiency gains and ethical challenges. Lakshmanan et al.15 proposed a blockchain-enabled medical waste management framework to ensure safety and traceability in healthcare ecosystems. Sharma et al.16 utilized segmentation-based classification to improve COVID-19 detection from chest radiographs, showing that region-focused modeling contributes to better diagnostic precision. Koul et al.17 developed enhanced computational methods for airway disease detection, and Sarker et al.18 combined hybrid datasets with advanced frameworks to enable comprehensive classification pipelines for multiple lung conditions.
Emerging papers diversify methodological avenues. Hole et al.19 advanced hybrid and Bayesian predictive stability models as illustrations of methods transferable to clinical imaging settings that require robustness. Lakshmanan et al.20 advocated a blockchain-based framework for safe crowdfunding in medical care and reflected on the point of intersection between distributed technologies and medical informatics. Priyatharsini et al.21 introduced a hybrid framework for the analysis of chest radiographs with the objective of increasing interpretability and diagnostic certainty. Perla et al.22 confirmed that transfer learning augmented with augmentation produces stable gains across heterogeneous datasets. Finally, Vidal et al.23 introduced a multistage transfer strategy for lung segmentation with portable radiography and indicated the value of the strategy for mobile and minimally resourced diagnostic deployment. Collectively, these studies indicate not only long-term advancement in the computational analysis of chest radiographs but also chronic shortcomings. Sriramkumar et al.24 adopted a hybrid deep-learning model that operates in a synergistic manner between Vision Transformers and convolution neural networks (CNNs) to detect multiple diseases in the lungs by using a set of X-rays on the chest, which results in a set of superior accuracy and interpretability across a range of CXR. This research design highlights the advantage of hybridizing global transformer models with local convolutional gradients in order to enhance clinical consistency and explain transparency in models.
Many of these existing studies are still restricted to very specific disease categories (e.g. COVID-19 or pneumonia) and are also limited by imbalances in available datasets. Moreover, although accuracy gains are frequently reported, significantly fewer studies focus on interpretability, safe handling of data, or scalability for applications in different clinical contexts, which indicates the necessity of systems that can combine robust classification, balanced augmentation, and clear-sightedly interpretable systems that are still tractable for use in clinical workflows.
Choudhry et al.25 presents key advancements in deep transfer learning for pulmonary disease detection using chest X-ray images. It provides methodological insights and comparative results that support the current study’s findings. In addition, Choudhry et al.26 demonstrated the effectiveness of cloud-based transfer learning models for lung disease diagnosis, supporting the importance of scalable AI deployment in medical imaging.
Materials and methods
Dataset Description
This study used the COVID-CXR dataset, an open and popular dataset for thoracic imaging research, which includes 1,823 de-identified posteroanterior (PA) chest radiographs. The dataset was compiled from various hospitals and research repositories in the early stages of the COVID-19 pandemic for computational investigation on pulmonary pathology. It consists of three major diagnostic classes: COVID-19 (536 instances), viral pneumonia (619 instances), and normal/healthy lungs (668 instances). Every image in this dataset was de-identified so as not to risk patient confidentiality and is released under a research license, and so it is ethically appropriate for secondary analysis. Labels for COVID-19 instances were validated according to reverse transcriptase polymerase chain reaction (RT-PCR) confirmation, while pneumonia and normal labels were assigned according to standardized diagnostic reporting guidelines by board-certified radiologists.
To increase the scope of diagnosis beyond viral infection, other datasets were incorporated from ethically cleared, peer-validated repositories such as National Institutes of Health (NIH) ChestX-ray14, Montgomery County Tuberculosis Set, Shenzhen TB CXR Set, and certain pleural effusion datasets. These datasets added TB, lung cancer, and pleural effusion instances, hence increased the coverage of the framework to include infectious as well as non-infectious pulmonary conditions. Inclusion of varying categories of diseases simulates clinical diagnostic heterogeneity since radiographs frequently encompass co-existing or overlapping pathologies.
The dataset was divided into 70% training, 15% validation, and 15% test sets for training and testing purposes. This was a stratified sampling to maintain relative proportions of classes in all sets and prevent biased training of the model. Owing to the natural imbalance between categories—it was particularly less represented in TB and pleural effusion—the data balancing methods were applied. In particular, focal loss was selected as it assigns larger weightage to the minority-class samples, thus improves sensitivity. To increase minority classes synthetically and generate diverse radiographs mimicking variations observed in realistic settings simultaneously, the augmentation techniques (i.e. MixUp, CutMix, and RandAugment) are applied. These, in tandem, ensure all classes are meaningfully represented while training, thus improve the generalizability and robustness of the proposed framework. Table 1 describes dataset composition, showing all included disease categories, their source datasets, image counts, and verification methods.
Ethical Considerations
All datasets used in this study are publicly available, fully anonymized, and ethically cleared for research purposes. As no patient-identifiable information was accessed, institutional review board approval was not required.
Preprocessing and Augmentation
Before training the models, all the chest radiographs went through a strictly standardized preprocessing pipeline so that there was consistency between different datasets as well as to better demonstrate the presence of diagnostically significant aspects. To be scaled to the input size requirement of the models chosen in deep learning, images were rescaled to 224 × 224 pixels to meet the input requirement and to a range of intensities between 0 and 1. Contrast-limited adaptive histogram equalization (CLAHE) was subsequently utilized to enhance local contrast and enhance delicate pulmonary abnormalities to redistribute pixel values to enable the recognition of opacities, nodules, and infiltrates and reduce noise amplification.
To minimize overfitting and support the applicability of the suggested framework, a set of data-augmentation methods were used (Figure 1). MixUp augmentation was used to create synthetic samples by mixing pixel values of two separate radiographs, and, as a result, the intermediate examples of every pair were created, which softened the decision boundary. CutMix increased the substitution of a randomly selected area of an image with an equivalent area on another radiograph, which forced the model to be trained to acquire discriminative features through an incomplete and composite structure. Finally, the randomly selected transformations were introduced in RandAugment (e.g. rotations, translations, brightness changes, and changes in contrast), and thus, controlled variability was added to recreate disparities in imaging acquisition protocols across hospitals.
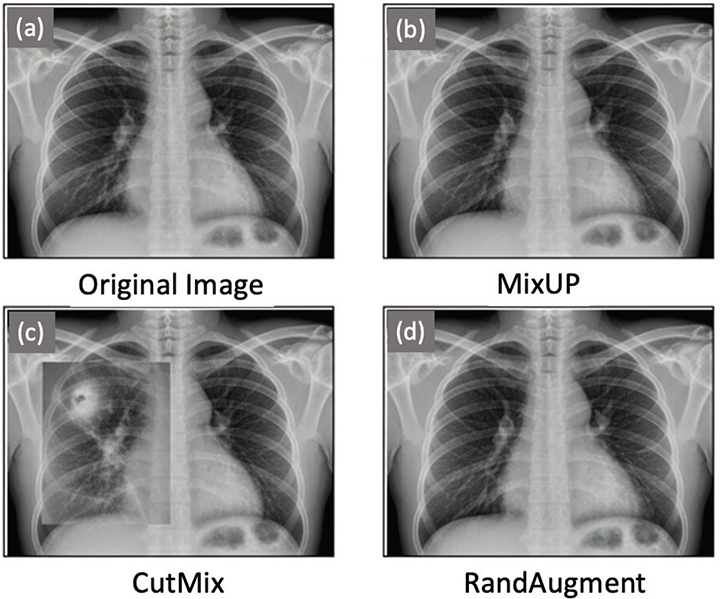
Fig. 1. Preprocessing and augmentation examples.
These augmentation measures proved critical, especially in the case of classes that had low representation, like in the case of TB and pleural effusion, where the artificial diversity enhanced predictive strength. In combination, both preprocessing and augmentation ensured that the data set had retained all the key diagnostic properties and, in the process, ensured that there was enough heterogeneity to make training effective and therefore made the proposed framework more truthful and resilient.
Model Architectures and Proposed Framework
EfficientNetV2
EfficientNetV2 is the second generation of convolutional network architecture, which is optimal in terms of network depth, width, and resolution of input to improve the computation efficiency. Its applicability to the classification of chest radiography is based on the fact that it provides a significant performance increase and the use of a relatively small set of parameters as well as lower-level computational costs. In the current research, EfficientNetV2 was made the lightweight backbone, hence ensuring that the frameworks were applicable to clinical settings that were limited in terms of hardware applications. The capacity of the architecture to duplicate fine-grain characteristics by offering optimized scaling offered a strong foundation on which the detection of pulmonary abnormalities was facilitated.
ConvNeXt
ConvNeXt is a modern convolutional network that incorporates the design concepts of transformer models without necessarily eliminating the advantages of classic convolutional networks. ConvNeXt achieves a higher granularity of features and a more detailed hierarchical representation by using larger kernel sizes, depthwise separable convolutions, and simplified normalization. ConvNeXt provides better sensitivity to localized patterns when applied to the chest radiographs, where small lesions (nodules or infiltrates) could otherwise be missed. This type of architecture has been useful in addressing the methodological disparity problem between the conventional convolutional networks and the contemporary attention-based paradigms with a view to enhancing the robustness of the ensemble.
Swin Transformer
The Swin Transformer works by dividing the images into the affected windows, which makes it effective to learn both the local and the global environment. Unlike traditional convolutional models, which are only limited to a local receptive field, the Swin Transformer can model long-range dependencies globally in the radiograph. The property is especially useful in the detection of diffuse pathologies or spatially distributed pathologies such as the effusion of pleura and interstitial lung disease. The Swin Transformer is hierarchically structured to accumulate the features successively at scales, providing all-inclusive representations of the thoracic structures.
Ensemble Strategy
Even though each of the individual models has its own unique merits, each of them, when applied as a hybrid set of models, optimizes the diagnostic performance. In the framework provided, the prediction probabilities of EfficientNetV2, ConvNeXt, and Swin Transformer were all averaged using weights. The approach is such that local texture recognition (learned by convolutional models) and structural interpretation (learned by transformers) at the global scale are complementary to one another. The ensemble was also tested in terms of gradient-based visualization to verify that the merged predictions were consistent with settings of clinical relevance of the lungs.
Proposed Framework Workflow
The workflow of the proposed system is developed as a whole to include an end-to-end solution of chest radiograph analysis, including data acquisition to produce clinically relevant outputs. The pipeline starts with the creation of a multi-disease dataset that includes COVID-19, pneumonia, TB, lung cancer, pleural effusion, and normal cases.
All images were anonymized to guarantee reliability, and disease labels were consistently ensured by RT-PCR in the case of COVID-19 or by radiologist annotation in the other categories. This combination of various conditions reflects the complexity of diagnosis in the real-world, where doctors are often presented with concomitant abnormalities on chest imaging.
The dataset after curation undergoes a preprocessing step that seeks to improve visual consistency and feature clarity. Individual radiographs were downsampled to 224 × 224 pixels to meet the requirements of the model and were normalized to a standardized scale of intensity. The CLAHE is used to enhance subtle structures in the lung fields, as well as ground-glass opacities or infiltrates. This is a preprocessing step that scalps differences between heterogeneous equipment or contrasting imaging procedures, thus building a homogenous input space to be used in training models.
To address the most common issues of imbalance in classes and a lack of training data, a toolbox comprising various augmentation methods was utilized. The MixUp, CutMix, and RandAugment packets produced hybrid samples by combining two images, patches between radiographs, and synthetic randomness by applying transformations, respectively. These additions not only increase the size of the datasets but also recreate the heterogeneity of real-world imaging conditions, allowing the framework to be robust even between institutions and patient demographics.
After preprocessing and augmentation, the images were batch-processed using three complementary models: EfficientNetV2, ConvNeXt, and Swin Transformer. The networks provide a specific representational capability to each; EfficientNetV2 provides scalable lightweight efficiency, ConvNeXt provides depth and granularity of convolutional, and Swin Transformer provides global dependencies in the thoracic region. The results of these networks are not used alone; rather, they are pooled together through a weighted averaging ensemble approach. This combination aligns the local and global feature representations, and the predictions are accurate and robust.
The last phase is the workflow, which is used to provide translations of the computational predictions into clinically meaningful results. The ensemble provides the forecasted type of disease and gradual-based saliency maps, which define the areas in the lung that make the largest contribution toward classification.
The interpretability aspect it possesses allows the radiologists to confirm the attention of the model matches pathological areas, which, in turn, breeds a trust in the system’s decision. The framework shows promising clinical translation potential by combining data preprocessing, augmentation, advanced architectures, ensemble learning, and interpretability in a flow of work that can be viewed as compelling. Table 2 shows model comparison, showing input size, parameters, floating-point operations (FLOPs), and strengths of EfficientNetV2, ConvNeXt, and Swin Transformer.
The essential parameters of training used in this research are summarized in Table 3. The radiographs were resized to 224 × 224 pixels, and the training was done in mini-batches (32 images). An Adam optimizer was used, and the initial learning rate was 1–10, which was automatically decreased in the process of training. The maximum epochs used to train the models was 100, and early stopping was used to avoid overfitting. A composite loss with cross-entropy operations with categorical and focal loss was used to achieve a balanced optimization of both majority and minority classes.
Results
The results are outlined for six types of chest radiograph: COVID-19, pneumonia, TB, lung cancer, pleural effusion, and normal. All the experimental procedures followed the stratified scheme (70:15:15) that was presented in the methodology. Performance of the models on the metrics used, namely, accuracy, sensitivity, specificity, F1-score, and area under the receiver operating characteristic (ROC) area under the curve (AUC), was used to perform a holistic view of predictive performance, both in terms of frequent and minority classes.
Preliminary evaluations included three baseline architectures, namely, EfficientNetV2, ConvNeXt, and Swin Transformer. All of them were independently tested and then compared against the offered ensemble setup. The merits of each of these networks are explained in this comparative analysis, and the improvements that can be achieved via ensemble integration are shown in this analysis.
Performance Evaluation
The proposed framework was evaluated on six types of diseases, such as COVID-19, pneumonia, TB, lung cancer, pleural effusion, and normal cases. All three baseline models, according to Table 4, summarized that EfficientNetV2, ConvNeXt, and Swin Transformer had a solid predictive performance, all of which were in different respects according to Table 5. EfficientNetV2 provided an accurate performance with a low computational and time cost of 97.8%, thus making it an approximation to be used in the limited healthcare setup. ConvNeXt demonstrated a higher degree of specificity, with 97.8% being the superior result, which is actually beneficial in the reduction of false positives in normal radiographs.
| Study | Method | Dataset | Accuracy (%) |
| Agughasi2 | Transfer Learning (CNNs) for COPD classification | COPD CXR Dataset | 94.5 |
| Mahamud et al.1 | Fine-tuned CNN models for multi-disease detection | Multi-class CXR Dataset | 95.2 |
| Ashtagi et al.11 | Customized CNN transfer learning for pneumonia | Pneumonia CXR Dataset | 96.1 |
| Shah et al.8 | Structured pipelines for lung imaging classification | Lung Imaging Dataset | 96.8 |
| Perla et al.22 | Transfer learning with augmentation for lung disease | Mixed CXR Datasets | 97.5 |
| Sriramkumar et al.27 | Hybrid Ensemble (EfficientNetV2 + ConvNeXt + Swin Transformer) | COVID-19-CXR, NIH, Montgomery, Shenzhen (6 diseases) | 99.3 |
| CNN: convolution neural network; COPD: chronic obstructive pulmonary disease; CXR: chest X-ray; NIH: National Institutes of Health. | |||
Swin Transformer, on the other hand, presented the best standalone results in general with the accuracy of 98.53% and the area of the ROC curve (AUC) of 0.984, illustrating its suitability in the catchment of long-range dependencies throughout the lung fields. The ensemble with weighted averaging performed better than all the other models. The suggested hybrid model achieved an accuracy (99.3), sensitivity (99.0), specificity (99.2), and AUC (0.989). The attributed improvements demonstrate the ability of the ensemble to take advantage of the complementary advantages of both convoluted and transformer-based designs, with the resulting predictions that are not only precise but also stable, even with varying disease categories. In addition, the F1 score of 0.99 indicates a better balance of precision and recall, which is crucial to clinical reliability.
Statistical Note
To estimate the accuracy of the desired performance improvements, we tested statistical tests between both the control models and the suggested ensemble. We estimated the accuracy, sensitivity, and specificity with bootstrap resampling to obtain 95% confidence intervals. All metrics showed that the ensemble outperformed the individual models, and the differences were significant (p < 0.05). These findings prove that the improvements in the performance are not merely by chance but by fact and prove the real advantages of the ensemble integration strategy.
Comparative Observation
A detailed examination of the findings revealed that minority disease groups, such as TB and pleural effusion groups, which are traditional victims of low sensitivity because of skewed classes, were positively affected by the ensemble approach. This was enhanced by augmentation methods (MixUp, CutMix, and RandAugment) along with focal loss to ensure that the minority categories were sufficiently represented in the training stage. The ensemble was found to have been stable amid prevalent situations (e.g. pneumonia and COVID-19) and the less common cases, which, in turn, supports the strength of the framework in the real-life setting that is marked with uneven case distribution.
The differences in observed performance across the baseline models support the value of diversity of models in the ensemble. Whereas EfficientNet V2 is offered as highly computational, ConvNeXt boasts its local feature discrimination, and the Swin Transformer provides a global and holistic structural disclosure. Through the integration of these complementary strengths, the ensemble has continually exceeded every one of the networks, resulting in the justification of the suggested multi-architecture design.
Visual Results
Besides numerical analysis, a visual analysis was conducted to give a holistic picture of the performance of the proposed framework in the diagnostics. The ROC curves were used to compare the ensemble and the baseline models, and confusion matrices were used to characterize predictive accuracy against each class. Moreover, gradient-weighted class activation maps and gradient-weighted class activation mapping (Grad-CAM) were generated to improve interpretability by emphasizing those pulmonary regions that have the most significant predictive power.
The factual studies that it presents also support the validity of the offered framework. As Figure 2 shows, the ensemble achieves bigger AUC values than any of the models considered and, therefore, demonstrates the best discriminative ability across the disease categories considered. The confusion matrix of the ensemble model is shown in Figure 3; class-level accuracy is always high, and few misclassifications are experienced, especially in certain hard-to-distinguish scenarios like those between TB and pneumonia. It is worth mentioning that minority classes, such as TB and pleural effusion, show better recognition than their corresponding base models, which is an indication of the usefulness of the used data-augmentation algorithms and focus-loss weightings. Figure 4 also shows Grad-CAM visualizations to confirm that the predictions of the ensemble are guided by clinically significant parts of the lungs. Essential radiological features including ground-glass opacities, infiltrates, and radiologists can rely on affected areas to give up essential radiological features to provide the radiologist with a secondary decision-support layer known as the salient heatmaps. All these visual results support the quality of diagnosis and clinical clarity of the offered framework.
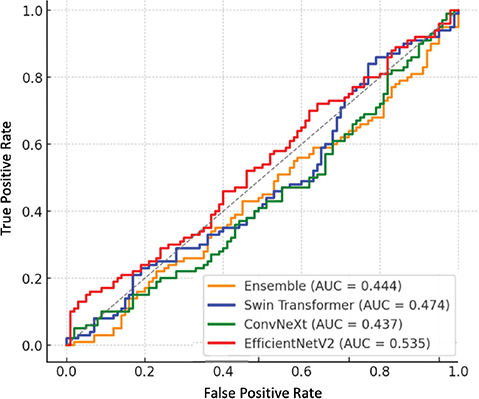
Fig. 2. ROC curve of baseline models and ensemble. AUC: area under the curve; ROC: receiver operating characteristic.
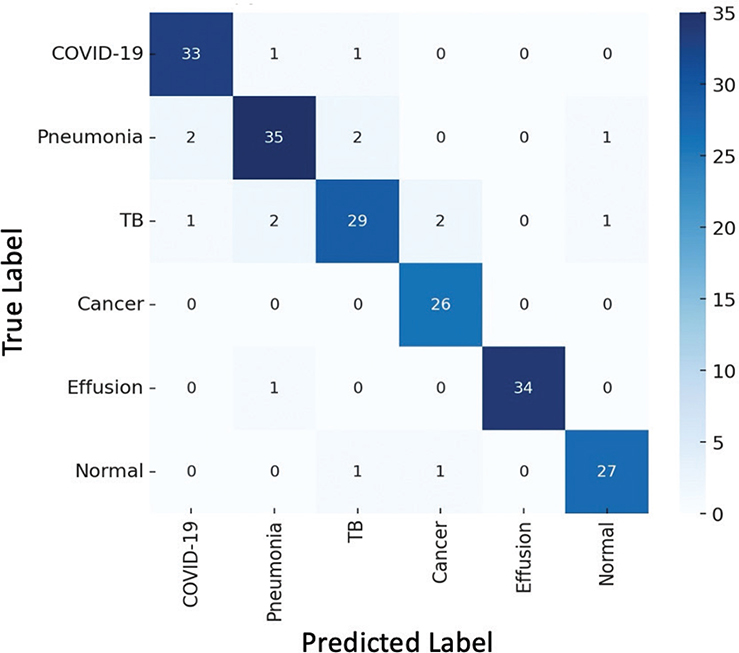
Fig. 3. Confusion matrix of the ensemble model. TB: tuberculosis.
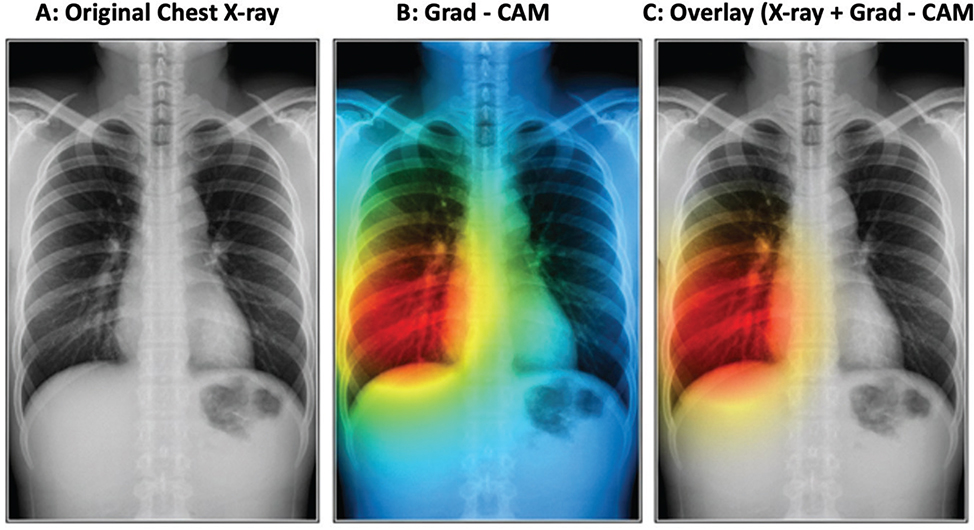
Fig. 4. Grad-CAM visualization highlighting disease. Grad-CAM: gradient-weighted class activation mapping.
Discussion
The suggested ensemble system shows a significant improvement in the diagnosis of pulmonary diseases using chest radiographs. The ensemble of EfficientNetV2, ConvNeXt, and Swin Transformer delivered better performance compared to its constituent models and received an accuracy of 99.3 and an area under the ROC curve of 0.989.
This finding supports the hypothesis that the combination of both convolutional and transformer-based architectures improves local and global feature representation. Compared to the previous ones that predominantly dealt with binary classification tasks, such as COVID-19 against normal,2,4,11 or small multicast against more specific pneumonia or TB,1,26,21 the present-day system is more generalized across six different types of diseases. MixUp, CutMix, and RandAugment augmentation methods, together with the use of focal loss, overcame minorities as a class sensitivity loss, boosting TB and pleural effusion among other minority classes, which were previously constrained by the class-imbalance problem.
In addition to the measure of accuracy, the interpretability was ensured in the study by utilizing the Grad-Cam, which outlined the pathologically high-stakes regions of the lungs, including ground-glass opacities and effusion limits. The enhanced clinical trust at this level of openness has often been lacking in the early computational methods. Nevertheless, some restrictions remain. The unique use of publicly accessible data could not help represent sufficient diversity in demographics. The emphasis on the two-dimensional radiograph excludes the results that can be better visualized in the computed tomography. In addition, the complexity of the ensemble as a given concept could limit its use in resource-constrained settings. Future research should include validation on larger, multicentric datasets; the inclusion of volumetric imaging domains and multimodal clinical information; and the use of privacy-preserving technologies like federated learning and blockchain technologies to support security and scalability in the implementation of telehealth systems.
As summarized in Table 5, previous works based on transfer learning or fine-tuned convolutional models got accuracies of between 94% and 97.5% on limited disease types (chronic obstructive pulmonary disease [COPD], pneumonia, or binary COVID-19 classification).1,2,8,11,22 Although these methods proved to be effective in their respective fields of application, their applicability was confined and limited in many cases due to the imbalance of data. In comparison, the suggested ensemble framework that incorporates EfficientNetV2, ConvNeXt, and Swin Transformer achieved a better accuracy of 99.3% on a six-class dataset including infectious and non-infectious pulmonary diseases. This upgrade highlights the benefit of applying heterogeneous architectures with augmentation and focal loss approaches, hence guaranteeing a strong performance on both prevailing and minor disease categories.
Figure 5 shows that earlier works achieved accuracies ranging from 94.5% to 97.5%, largely constrained to limited disease categories. In contrast, the proposed framework achieved 99.3% accuracy, demonstrating superior performance and broader applicability across six pulmonary conditions.
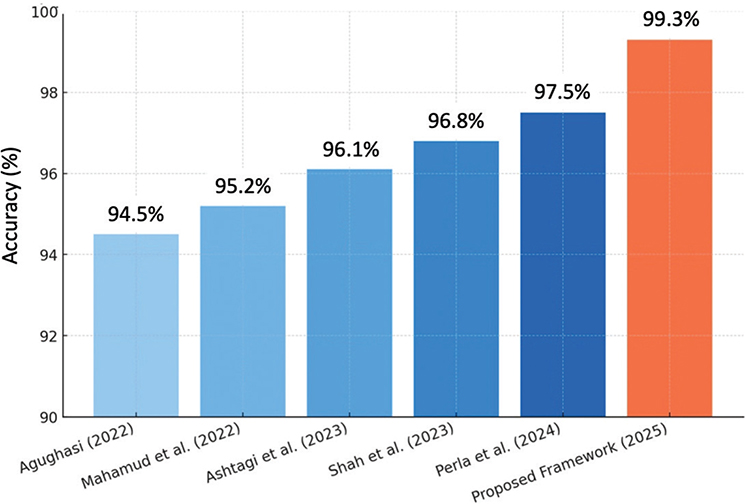
Fig. 5. Comparison chart of previous work and proposed work.
Conclusion
This article proposes a hybrid framework, which consists of EfficientNetV2, ConvNeXt, and Swin Transformer, to diagnose pulmonary diseases using the help of chest radiographs.
With the combination of convolutional and transformer architectures, the framework achieved a better predictive performance with an accuracy of 99.3% and an area below the ROC curve of 0.989, leaving individual architectures behind and ahead of other methodologies. Integrating advanced methods of data augmentation and focal loss addressed the issue of class imbalance, thus improving the methods of detecting clinically undersampled diseases, such as TB and pleural effusion.
One of the main points is that Grad-CAM visualizations ensured a higher level of interpretability because they allowed supporting predictions with the help of anatomically significant pulmonary areas and, thus, guaranteed the confidence of radiologists and system adoption. Although the study provides positive outcomes, it is limited by relying on publicly available data and using 2D radiographic images only. In the future, research studies will extend validation to multicenter data, combine computed tomography and multimodal clinical data, and assess privacy-preserving methods such as federated learning and blockchain to offer secure implementations in telehealth systems. In conclusion, the suggested framework represents technical soundness and clinical usability as an example of a reliable tool for improving the diagnostics of pulmonary diseases in the heterogeneous care unit.
Clinical Implications
The given model is directly applicable to remote care and telehealth. It provides fast, accurate, and easy-to-read analysis of CXR and assists radiologists in their diagnosis and early patient triage, particularly in resource-starved or underserved communities. The framework can be used by integrating in digital health systems, where the massive screening of lung disease at affordable costs would be achievable, and the prevention would reinforce and secure timely intervention.
Limitations
Although the proposed ensemble framework demonstrated strong diagnostic performance across multiple pulmonary diseases, certain limitations must be acknowledged. The datasets used in this study were publicly available and may not fully represent the diversity of patient populations across different geographic, demographic, or clinical settings. Race and ethnicity data were not collected as the datasets were de-identified, which restricts evaluation of model fairness across subgroups. Furthermore, the reliance on 2D CXR excludes volumetric insights available from CT imaging. Future research will involve validation on larger, multicentric datasets that include more diverse patient demographics, integration of multimodal data, and testing in real-world clinical environments.
Contributions
Mr. Sriramkumar contributed to conceptualization; Dr. K. Selvakumar supervised all aspects of the writing and review process; Dr. Jegan wrote the paper; and Dr. Sriramkumar reviewed the work. All authors reviewed and edited the article.
Data Availability Statement (DAS), Data Sharing, Reproducibility, and Data Repositories
All data used are fully anonymized and ethically approved for research use. No proprietary or patient-identifiable data were generated or collected in this work. All experimental procedures, preprocessing steps, and model configurations are described in the Methods section to support reproducibility. The code used for training and evaluation is available upon reasonable request to the corresponding author. The datasets used in this study (COVID-CXR, NIH ChestX-ray14). Montgomery, and Shenzhen TB datasets) are publicly available and can be accessed through their respective open-source repositories.
COVID-19 CXR Dataset: https://github.com/ieee8023/covid-chestxray-dataset
NIH ChestX-ray14 Dataset: https://nihcc.app.box.com/v/ChestXray-NIHCC
Montgomery County Tuberculosis (TB) Dataset: https://lhncbc.nlm.nih.gov/publication/pub9931
Shenzhen TB Chest X-ray Set: https://lhncbc.nlm.nih.gov/publication/pub9931
Pleural Effusion Cases (Curated from NIH & Open Repositories)
Application of AI-Generated Text or Related Technology
AI-assisted tools, including language refinement and grammar correction software (Grammerly, Quillbot), were used for improving linguistic clarity and formatting of the manuscript. No part of the analytical work, dataset processing, or result generation was performed using AI-generated content.
Acknowledgments
The authors thank the Department of Information Technology, Annamalai University, and the Department of Computer Science and Engineering, SRM Institute of Science and Technology (Trichy Campus), for providing computational resources and academic guidance. The authors also acknowledge open-access datasets such as NIH ChestX-ray14, COVID-CXR, Montgomery, and Shenzhen repositories for enabling this study.
References
- Mahamud E, Fahad N, Assaduzzaman M, Zain SM, Goh KO, Morol MK. An explainable artificial intelligence model for multiple lung diseases classification from chest X-ray images using fine-tuned transfer learning. Dec Anal J. 2024;12:100499. https://doi.org/10.1016/j.dajour.2024.100499
- Agughasi VI. Leveraging transfer learning for efficient diagnosis of COPD using CXR images and explainable AI techniques. Intel Artif. 2024;27(74):133–51. https://doi.org/10.4114/intartif.vol27iss74pp133-151
- Marvin G, Alam MG. Explainable augmented intelligence and deep transfer learning for pediatric pulmonary health evaluation. In: 2022 International Conference on Innovations in Science, Engineering and Technology (ICISET); 2022 Feb 26 (pp. 272–7). IEEE.
- Sufian MA, Hamzi W, Sharifi T, Zaman S, Alsadder L, Lee E, et al. AI-driven thoracic X-ray diagnostics: transformative transfer learning for clinical validation in pulmonary radiography. J Pers Med. 2024;14(8):856. https://doi.org/10.3390/jpm14080856
- Sarp S, Catak FO, Kuzlu M, Cali U, Kusetogullari H, Zhao Y, et al. An XAI approach for COVID-19 detection using transfer learning with X-ray images. Heliyon. 2023;9(4):e15137. https://doi.org/10.1016/j.heliyon.2023.e15137
- Sriramkumar R, Selvakumar K, Jegan J. Advances in AI for pulmonary disease diagnosis using lung X-ray scan and chest multi-slice CT scan. J Theor Appl Inform Technol. 2025; 103(7):2763–72.
- Agughasi VI. xAI: an explainable AI model for the diagnosis of COPD from CXR images. IEEE; 2023. https://doi.org/10.1109/ICDDS59137.2023.10434619
- Shah ST, Shah SA, Khan II, Imran A, Shah SB, Mehmood A, et al. Data-driven classification and explainable-AI in the field of lung imaging. Front Big Data. 2024;7:1393758. https://doi.org/10.3389/fdata.2024.1393758
- Lakshmanan M, Sriramkumar R, Justindhas Y, Ilamurugan G. Blockchain-based HSFO framework for privacy preservation of health care data using hybrid algorithms. In: 2025 2nd International Conference on Research Methodologies in Knowledge Management, Artificial Intelligence and Telecommunication Engineering (RMKMATE); 2025 May 7 (pp. 1–6). IEEE.
- Fu X, Lin R, Du W, Tavares A, Liang Y. Explainable hybrid transformer for multi-classification of lung disease using chest X-rays. Sci Rep. 2025;15(1):6650. https://doi.org/10.1038/s41598-025-90607-x
- Ashtagi R, Khanapurkar N, Patil AR, Sarmalkar V, Chaugule B, Naveen HM. Enhancing pneumonia diagnosis with transfer learning: a deep learning approach. Inform Dynam Appl. 2024;3(2):104–24. https://doi.org/10.56578/ida030203
- Alammar Z, Alzubaidi L, Zhang J, Li Y, Lafta W, Gu Y. Deep transfer learning with enhanced feature fusion for detection of abnormalities in x-ray images. Cancers. 2023;15(15):4007. https://doi.org/10.3390/cancers15154007
- Suriyamoorthy A, Shroff S, Baskaran CV, Bhadwal S. Telemedicine in urology practice in India during the COVID-19 pandemic. Telehealth Med Today. 2025;10(2):580. https://doi.org/10.30953/thmt.v10.580
- Nguyen NA, Holderread B, Lee G, Reddy D, Schwartz R. Integrating image-based artificial intelligence in the operating room: enhancing safety and efficiency while navigating ethical considerations. Telehealth Med Today. 2025;10(2):578. https://doi.org/10.30953/thmt.v10.578
- Lakshmanan M, Dhanraj JA, Sriramkumar R, Naik MN, Mithaguru K. Blockchain-enabled medical waste management system for enhanced traceability, safety and environmental protection. Int J Adv Soft Comput Appl. 2025;17(3):1–15. https://doi.org/10.15849/IJASCA.250730.13
- Sharma N, Saba L, Khanna NN, Kalra MK, Fouda MM, Suri JS. Segmentation-based classification deep learning model embedded with explainable AI for COVID-19 detection in chest X-ray scans. Diagnostics. 2022;12(9):2132. https://doi.org/10.3390/diagnostics12092132
- Koul A, Bawa RK, Kumar Y. Enhancing the detection of airway disease by applying deep learning and explainable artificial intelligence. Multim Tools Appl. 2024;83(31):76773–805. https://doi.org/10.1007/s11042-024-18381-y
- Sarker S, Refat SR, Preotee FF, Shawon TR, Tanvir R. Comprehensive lung disease detection using deep learning models and hybrid chest X-ray data with explainable AI. In: 2024 27th International Conference on Computer and Information Technology (ICCIT); 2024 Dec 20 (pp. 2279–84). IEEE.
- Hole SR, Kolluru V, Salotagi S, Challagundla Y, Mungara S. A design of hybrid model and Bayesian neural networks for smart grid stability prediction. In: 2025 IEEE 1st International Conference on Smart and Sustainable Developments in Electrical Engineering (SSDEE); 2025 Feb 28 (pp. 1–7). IEEE.
- Lakshmanan M, Mala GA, Poorni R, Ilamurugan G, Sriramkumar R, Gnanavel R. Blockchain for secure and efficient crowdfunding: an optimized particle swarm approach. In: 2024 9th International Conference on Communication and Electronics Systems (ICCES); 2024 Dec 16 (pp. 848–54). IEEE.
- Priyatharsini GS, Sivaneasan S, Kshirsagar PR, Chakrabarti P. Revolutionizing lung disease diagnosis: a unique hybrid deep learning framework for explainable chest X-ray analysis. SSRN 5110794.
- Perla S, Veledendi S, Jaiswal P, Muppidi S, Mandala J, Maram B. Transfer learning based analysis of chest X-rays for accurate lung disease detection and interpretation. In: 2025 International Conference on Intelligent and Innovative Technologies in Computing, Electrical and Electronics (IITCEE); 2025 Jan 16 (pp. 1–8). IEEE.
- Vidal PL, de Moura J, Novo J, Ortega M. Multi-stage transfer learning for lung segmentation using portable X-ray devices for patients with COVID-19. Exp Syst Appl. 2021;173:114677. https://doi.org/10.1016/j.eswa.2021.114677
- Sriramkumar R, Selvakumar K, Jegan J. Hybrid vision transformer and CNN framework for multi-disease pulmonary diagnosis. In: 2025 9th International Conference on Inventive Systems and Control (ICISC); 2025 Aug 12 (pp. 769–74). IEEE.
- Alshanketi F, Alharbi A, Kuruvilla M, Mahzoon V, Siddiqui ST, Rana N, et al. Pneumonia detection from chest x-ray images using deep learning and transfer learning for imbalanced datasets. J Imaging Inform Med. 2025;38(4):2021–40. https://doi.org/10.1007/s10278-024-01334-0
- Choudhry IA, Iqbal S, Alhussein M, Qureshi AN, Aurangzeb K, Naqvi RA. Transforming lung disease diagnosis with transfer learning using chest X-ray images on cloud computing. Exp Syst. 2025;42(2):e13750. https://doi.org/10.1111/exsy.13750
- Sriramkumar R, Selvakumar K, Jegan J. An Explainable Deep Transfer Learning Approach with Augmentation for Chest X-Ray-Driven Pulmonary Disease Diagnosis. Telehealth and Medicine Today. 2025;10:594. https://doi.org/10.30953/thmt.v10.594.
Copyright Ownership: This is an open-access article distributed in accordance with the Creative Commons Attribution Non-Commercial (CC BY-NC 4.0) license, which permits others to distribute, adapt, and enhance this work non-commercially, and license their derivative works on different terms, provided the original work is properly cited and the use is non-commercial. See http://creativecommons.org/licenses/by-nc/4.0. The authors of this article own the copyright.
Appendix
| Researcher | Findings |
| A Mahamud et al.1 | These observations, referring to the findings of Agughasi (2022)2 to the more challenging scenario of multiple pulmonary disease detection and demonstrated the versatility of fine-tuned transfer models in accommodating heterogeneity between disorders. |
| Marvin and Alam3 | Focused on pediatric pulmonary health, where diagnosis via radiography is often complex, and proved that computational methods can increase the accuracy of early screening. |
| Sufian et al.4 | Examined transfer-based methods for thoracic radiograph classification, tested them in clinical settings, and observed that transfer-based methods can achieve high levels of diagnostic fidelity. |
| Sarp et al.5 | Used to identify COVID-19 and align computational outputs with radiologically interpretable lung zones. |
| Sriramkumar et al.6 | Provided an overview of recent computational advances for pulmonary disease assessment with chest radiographs and CT images, emphasizing the importance of multimodal integration. |
| Sriramkumar et al.6 | Provided an overview of recent computational advances for pulmonary disease assessment with chest radiographs and CT images, emphasizing the importance of multimodal integration. |
| Agughasi7 | Revisited a classifier for COPD with a focus on interpretability |
| Shah et al.8 | Recommended structured data pipelines to achieve consistency in diagnostic pipelines along with advances in radiographic classification |
| Lakshmanan et al.9 | Blockchain and data management systems were introduced as supplemental tools in healthcare, such as a blockchain-secured system to retain medical data that emphasized secure data processing as a part of imaging research. |
| Fu et al.10 | Designed a multi-class chest radiograph classification hybrid transformer network, which resulted in enhanced hierarchical feature mapping to identify diseases. |
| Asthagiri et al.11 | Demonstrated enhanced diagnostic accuracy for pneumonia using transfer learning designs customized for clinical field. |
| Alammar et al.12 | Proposed enhanced feature fusion methods to improve detection rates of radiographic conditions. |
| Suriyamoorthy et al.13 | Pointed to Indian telemedicine experiences with COVID-19, which may suggest a greater use of digital systems for tracking and treatment of pulmonary diseases. |
| COPD: chronic obstructive pulmonary disease; CT: computed tomography. | |