ORIGINAL RESEARCH
Early Prediction of Preeclampsia Using Ensemble Machine Learning With Multi-Method Explainable AI Approach
Sari Puspita, S.Si, M.Si  ; Gusrino Yanto, S.Kom, M.Kom
; Gusrino Yanto, S.Kom, M.Kom  ; Rifa Turaina, S.Kom, M.Kom
; Rifa Turaina, S.Kom, M.Kom  ; Nency Extise Putri, S.Kom, M.Kom
; Nency Extise Putri, S.Kom, M.Kom 
Program Study of Information System, Faculty of Information Technology and Creative Industries, Universitas Metamedia, Padang, West Sumatera, Indonesia
Keywords: ensemble of machine learning, LIME, multi-method XAI, preeclampsia, SHAP
Abstract
Preeclampsia is a pregnancy complication that endangers the mother and fetus. Early detection is necessary to prevent serious complications. Here, the authors employ mixed methods, which combine quantitative and qualitative approaches to design a preeclampsia risk prediction system for pregnant women. The system uses four machine learning algorithms: logistic regression, decision tree, support vector machine (SVM), and random forest (RF). We conducted the evaluation process by using a multi-method explainable AI (XAI) approach with Shapley additive explanations, local interpretable model-agnostic explanations, and permutation feature importance to enhance the transparency and ease of interpretation of the results. Clinical variables included systolic and diastolic blood pressure, blood glucose, body temperature, heart rate, age, and urine protein. The results show that RF and SVM achieved the highest accuracy (78%) with relatively stable performance across risk categories. Multi-method XAI analysis indicated that blood pressure and blood glucose frequently appeared among influential features, although their relative importance varied depending on the model and explainability method. However, due to the limited dataset size and use of internal validation only, these findings should be interpreted as preliminary and multi-method XAI early identification of preeclampsia risk factors, not to replace clinical diagnosis or function as a standalone clinical decision-making tool.
Plain Language Summary
Preeclampsia is a serious pregnancy complication that can endanger both mother and baby. Early risk identification is essential to support timely monitoring and care. This study analyzed anonymized data from 299 pregnant women at a community health center in Padang, Indonesia. Several machine learning models were used to classify preeclampsia risk into low, medium, and high categories. Random forest and support vector machine achieved the most consistent results, with 78% accuracy. Explainable artificial intelligence (XAI) methods were applied to clarify how predictions were made. Blood pressure and blood sugar frequently influenced the results. The proposed model is intended to support early risk screening and does not replace clinical diagnosis or medical judgment. Further validation is required before clinical implementation.
Citation: Telehealth and Medicine Today © 2026, 11: 663 - https://doi.org/10.30953/thmt.v11.663
DOI: https://doi.org/10.30953/thmt.v11.663
Copyright: © 2026 The Authors. This is an open-access article distributed in accordance with the Creative Commons Attribution Non-Commercial (CC BY-NC 4.0) license, which permits others to distribute, adapt, enhance this work non-commercially, and license their derivative works on different terms, provided the original work is properly cited and the use is non-commercial. See http://creativecommons.org/licenses/by-nc/4.0. The authors of this article own the copyright.
Received: November 25, 2025; Accepted: January 30, 2026; Published: March 31, 2026.
Corresponding Author: Sari Puspita, Email: saripuspita@metamedia.ac.id
Competing interests and funding: The authors declare no conflicts of interest.
The authors received financial support for publication from the Ministry of Higher Education, Science, and Technology.
Preeclampsia is a hypertensive disorder that occurs during pregnancy. It afflicts approximately 5% to 8% of high-risk pregnancies and is one of the leading causes of maternal and perinatal morbidity and mortality worldwide.1–4 Preeclampsia usually occurs after 20 weeks of gestation and is characterized by high blood pressure, which is often accompanied by proteinuria or signs of other organ damage.2,5,6
If undetected or untreated, preeclampsia can progress to eclampsia. Eclampsia is a more severe condition characterized by seizures and can lead to fatal complications for the mother and fetus.7,8 The high rates of maternal and infant mortality due to preeclampsia highlight the need for an early detection system that is both more accurate and more efficient.9–11 Today, identifying the risk of preeclampsia requires manual clinical and laboratory examinations, which are time-consuming and resource-intensive.12,13 Additionally, the numerous interrelated risk factors, including maternal age, history of hypertension, previous pregnancies, body mass index, and genetic conditions, make the diagnostic process more complex.14
Previous studies on predicting preeclampsia used an ensemble of machine learning (ML) algorithms.15–17 They demonstrated the high accuracy of models such as random forest (RF) and eXtreme Gradient Boosting.18–20 The main obstacle is the lack of transparency in the clinical field. Multi-method explainable AI (XAI), including Shapley additive explanations (SHAP), local interpretable model-agnostic explanations (LIME) and glass-box explainable boosting machine models, are used to explain variables, improve interpretability, and maintain accuracy. This supports model interpretability and transparency within clinical research contexts.21,22 Previous preeclampsia prediction studies largely relied on single explainability techniques with a focus primarily on predictive accuracy, limiting a comprehensive understanding of model behavior. By integrating multiple complementary multi-method XAI, this study addressed this limitation.22
Although several physiological variables, such as body temperature and heart rate, were included in the predictive models, these features are not well-established independent clinical risk factors for preeclampsia. In this study, these variables were considered supportive indicators of general maternal physiological conditions, rather than primary clinical determinants. Therefore, conclusions regarding their clinical relevance were drawn cautiously, and the predictive importance of these non-standard variables should not be extrapolated beyond the context of early risk screening.
In Padang City, Indonesia, there were approximately 17,376 pregnant women in a population of 942,938 in 2022.23,24 In the same year, the maternal mortality rate dropped to 17 cases per 100,000 live births from 30 cases in 2021. It then increased to 23 cases in 2023.24 Preeclampsia in pregnant women is one cause of maternal and infant mortality. Healthcare professionals aim to identify preeclampsia early to ensure the safety of mothers during pregnancy, childbirth, and postpartum. According to the operational definition, approximately 20% of pregnant women are considered high-risk, which includes those with preeclampsia.
Figure 1 shows that the detection rates among high-risk pregnant women in 2023 were 113.4%.25 The data on the detection rate of high-risk pregnant women show that artificial intelligence (AI) technology has the potential to support decision-making,26,27 especially in terms of predicting preeclampsia. One of the frequently used methods is ML, a branch of AI that allows computers to learn and make decisions or predictions without explicit programming. This study uses four ML algorithms—logistic regression, decision tree, support vector machine (SVM), and RF—to classify preeclampsia risk levels in pregnant women.
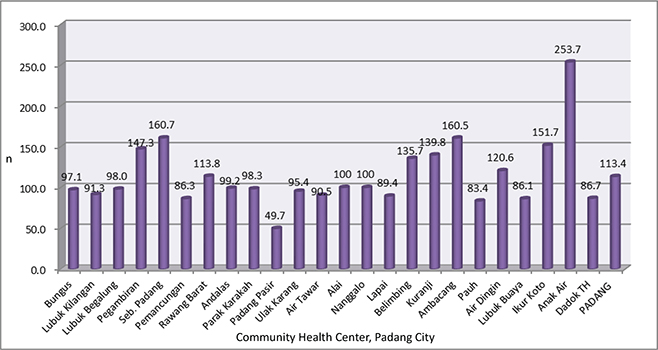
Fig. 1. Detection rates of high-risk pregnant women by health workers in 2023.25
Source: DINAS KESEHATAN KOTA PADANG [Internet], 2025.25
RF is an ensemble-based classifier that combines multiple decision trees to improve predictive stability.28–35 Logistic regression is a probabilistic linear classifier commonly used as a baseline model in medical prediction.32,36,37 A decision tree is a rule-based model that provides interpretable classification decisions,38–40 and an SVM is a margin-based classifier that is effective for handling high-dimensional clinical data.41–44
To ensure a transparent understanding of the prediction results from each model, this study applies a multi-method XAI approach,45,46 which helps explain and improve understanding of how models make decisions.47 In addition, three models were employed: SHAP, LIME, and permutation feature importance (PFI). SHAP uses principles of game theory to measure how much each feature contributes to the predicted result.48 Meanwhile, LIME is developing models to explain individual predictions and assess feature influence on performance using PFI.49–51 These techniques provide technical transparency regarding model behavior and feature contributions. However, the interpretability benefits are assessed at a methodological level and were not formally evaluated through clinician usability testing.
Methods
This study uses an ML modeling approach with a multi-method XAI method to predict preeclampsia risk in pregnant women. The modeling process is carried out in several systematic stages, as shown in Figure 2.
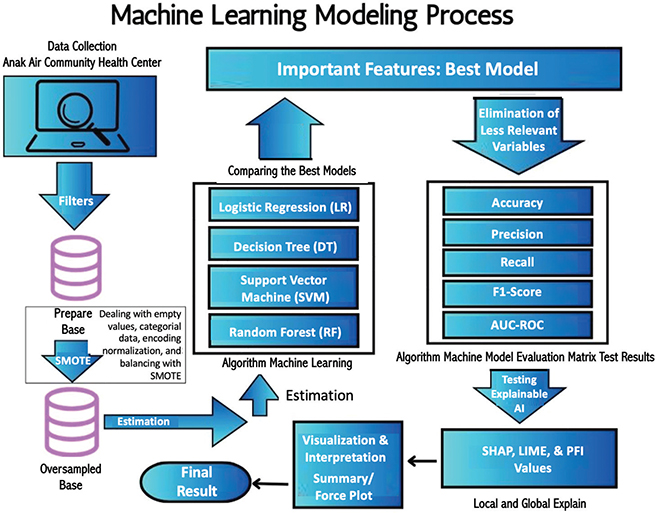
Fig. 2. Stages of the machine learning modeling process for preeclampsia risk prediction. AI: artificial intelligence; AUC ROC: area under the receiver operating characteristic curve; LIME: local Interpretable model-agnostic explanations; PFI: permutation feature importance; SHAP: Shapley additive explanations; SMOTE: synthetic minority oversampling technique.
There are the sequences of the preeclampsia prediction modeling process by using ML algorithms:
Data Collection
These collected data contain the medical information and clinical characteristics of pregnant women. The initial data undergo a filtering stage to remove incomplete, inconsistent, or irrelevant entries. This ensures that the data are high-quality, accurate, and suitable for effective and valid ML modeling analysis.
Prepare Base Data
The data preprocessing stage includes handling missing values, encoding categorical variables into numerical form, and normalizing continuous variables using min-max scaling to ensure comparable feature ranges. Outliers are retained to preserve clinically meaningful extreme values, and no domain-specific thresholds were applied in order to avoid introducing subjective bias. Class imbalance was addressed using the synthetic minority oversampling technique (SMOTE).52–54 SMOTE generates synthetic data for minority classes, balancing the dataset and preventing ML models from favoring the majority.55–57
Model with Machine Learning Algorithm
This study uses four ML algorithms—logistic regression, decision tree, SVM, and RF—to classify preeclampsia risk levels in pregnant women.58–60
Model Evaluation
Model performance: It is evaluated using quantitative metrics such as accuracy, precision, recall, and F1-score. These metrics evaluate the model’s capacity for precise predictions and its ability to balance the detection of positive and negative cases, especially in datasets with skewed class distributions. The model will be evaluated using the following metrics:
Accuracy: It measures how often the model makes correct predictions, which are compared to the total number of predictions.61,62
Precision: It measures how many positive predictions are correct, which are compared to all positive predictions made by the model.63,64
Recall (i.e., sensitivity or true positive rate): It measures how many positive examples are detected by the model compared to the total number of examples in the dataset.10
F1-Score: a metric that balances precision and recall by combining them into a single value. It is useful when there is an imbalanced class or when both false positives and false negatives must be considered simultaneously.65
Model evaluation was conducted using an internal validation framework. Due to dataset size constraints, nested cross-validation and external validation with an independent test set were not performed. Consequently, the reported performance metrics reflect the model’s internal behavior and should be interpreted as preliminary screening performance rather than definitive clinical accuracy. Formal statistical uncertainty estimation, such as confidence intervals for accuracy or area under the curve (AUC), and hypothesis-based statistical comparisons between models were not performed due to the limited dataset size and the exploratory screening nature of this study. In addition, model performance was not evaluated on an independent, untouched external test set. Therefore, the reported metrics should be interpreted as descriptive indicators of internal screening performance rather than statistically validated estimates of generalizable clinical accuracy. Accordingly, no claims of model superiority or clinical equivalence are made.
Importance and Elimination Feature: It analyzes the level of importance of each feature in its contribution to the prediction. Identifying these important features determines which variables influence the model output the most.66 Features with low importance values are considered less relevant and eliminated from the modeling process. This elimination simplifies the model structure, reduces computational complexity, and improves interpretability without sacrificing prediction performance.67
Application of multi-method XAI: To improve the transparency and interpretability of the prediction model, this study uses three methods, namely, SHAP, LIME, and PFI.48,68,69
Visualization and Interpretation
The results of the model interpretation process are presented through two types of plots: summary plots and force plots. Summary plots illustrate the contribution of features to the prediction results, while force plots provide specific explanations at the individual level. Summary plots offer an overall view of how each feature influences the model’s predictions, while force plots provide specific explanations at the individual level.48,68,69
Final Result
The final result of the estimation and interpretation process is compiled into comprehensive findings that support decision-making in medical contexts. This information is designed to help healthcare professionals detect the risk of preeclampsia earlier.6,22,34
Results
Data were collected using field research methods from pregnant women. Then, the data were processed in Google Collaboratory with the Python programming language.70,71
Dataset of Pregnant Women
This study has several limitations related to the size of the dataset. With 299 records, the dataset increases the risk of overfitting and limits the generalizability of the findings to broader clinical populations. Although SMOTE was applied to address class imbalance, the use of synthetic samples might introduce noise and potentially distort underlying clinical relationships, especially in small datasets. Consequently, the reported performance should be interpreted with caution. Future studies should prioritize larger, multicenter datasets and external validation to ensure model robustness and clinical reliability. Table 1 shows an overview of the data used.
| No. | Age (yrs) | Systolic BP (mmhg) | Diastolic BP (mmhg) | Blood sugar (mg/dL) | Body temp (°F) | Heart rate (BPM) | Urine protein (mg/dL) | Risk* |
| 0 | 34 | 130 | 80 | 15.0 | 98 | 86 | 109.99 | High |
| 1 | 34 | 140 | 90 | 121.0 | 98 | 70 | 117.97 | High |
| 2 | 29 | 90 | 70 | 118.0 | 100 | 80 | 147.26 | High |
| 3 | 33 | 140 | 85 | 7.0 | 98 | 70 | 112.28 | High |
| 4 | 40 | 120 | 60 | 6.1 | 98 | 76 | 197.08 | Low |
| *Risk stratification (low, medium, and high risk). The three-class risk stratifications (low, medium, and high risk) reflect operational screening categories used in routine antenatal care at the study site. These labels were derived from existing clinical records and were not independently validated by specialist clinicians. bpm: beats per minute, BP: blood pressure; F: temperature reported in the Fahrenheit scale; mg/dL: milligrams per deciliter; yrs: years. | ||||||||
To facilitate the processing of the classification algorithm, the risk level as a target variable is converted from the categories of low, medium, and high risk to the numerical values 0, 1, and 2, respectively. The three-class risk stratification (low, medium, and high risk) reflects operational screening categories used in routine antenatal care at the study site. These labels were derived from existing clinical records and were not independently validated by specialist clinicians; therefore, they should be interpreted as screening-level risk indicators rather than diagnostic classifications.
As shown in Figure 3, the distribution visualization illustrates the initial imbalance of the data before balancing. The highest number is in the low-risk category (0), then it is followed by the high-risk category (2), and the lowest number is in the medium-risk category (1).
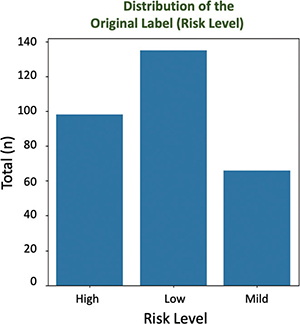
Fig. 3. The risk for preeclampsia.
Distribution of Preeclampsia Risk After SMOTE Implementation
The data distribution shows class imbalance, which affects model performance. This issue can be addressed through stratified sampling, oversampling, or weighting the minority class to improve the balance of risk prediction.
SMOTE application successfully balanced the amount of data in each risk label, which was previously uneven, eliminating bias toward the dominant class. Accordingly, each category (Label 0, Label 1, and Label 2) has 108 samples. This helps the model to recognize patterns evenly and improves the accuracy of risk predictions (Figure 4).
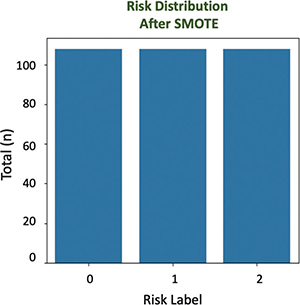
Fig. 4. Distribution of preeclampsia risk labels after SMOTE implementation. SMOTE: synthetic minority oversampling technique.
Confusion Matrix
A confusion matrix evaluates the performance of a classification model by showing the number of correct and incorrect predictions in each category. This facilitates analysis of the model’s accuracy and classification error.
Figure 5 shows that the classification performance varied among the models. The medium-risk group was the most challenging to predict. The RF and SVM models demonstrated the most balanced performance across all risk categories, whereas the decision tree and logistic regression models showed reduced sensitivity in the medium-risk class. These results suggest that ensemble and margin-based models are more effective for screening-oriented risk stratification. The comparison of model performance is shown in Table 2.
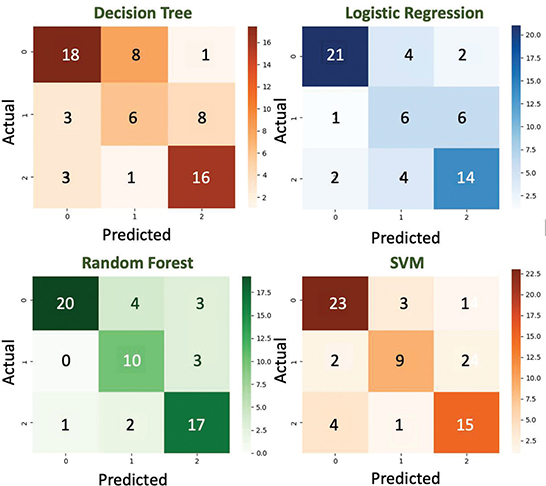
Fig. 5. Confusion matrix for four classification models. The y-axis is the number of the data based on risk level. SVM: support vector machine.
ROC Curve
The ROC curve evaluates a model’s ability to distinguish between classes. It displays the relationship between the true positive and false positive rates. Performance is measured by the AUC.
As shown in Figure 6, RF and SVM had the most consistent discriminative performance across all risk categories. Meanwhile, decision tree and logistic regression showed greater variability. Overall, ensemble-based models demonstrated superior class separability under the applied internal validation framework.
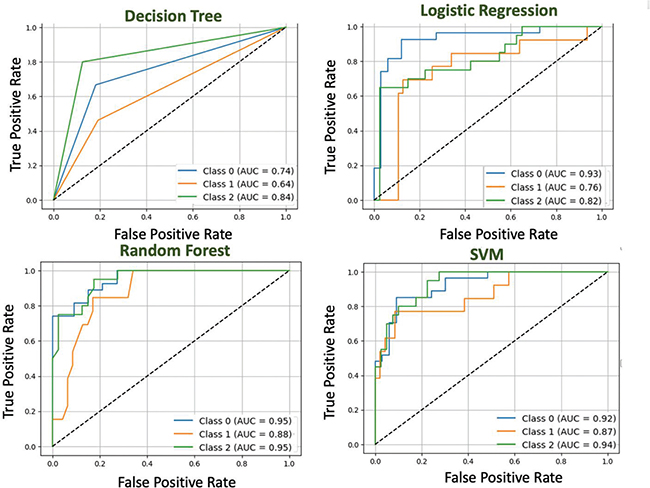
Fig. 6. Classification of receiver operating characteristic (ROC) curve for all evaluated models. AUC: area under the curve; SVM: support vector machine.
Visualization SHAP
SHAP is used to explain how each feature contributes to a model’s prediction. It provides a transparent, in-depth interpretation of the decisions that are made by an ML algorithm.
Figure 7 shows that the SHAP analysis revealed that dominant features varied across models, reflecting differences in learning mechanisms. Although their relative importance differed depending on the algorithm, blood pressure and blood glucose variables frequently appeared among the top contributors.
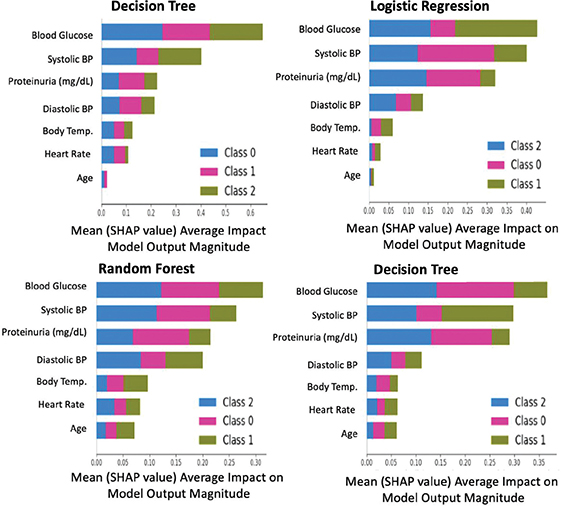
Fig. 7. SHAP visualization illustrating the contribution of each feature to the model’s prediction. BP: blood pressure; SHAP: Shapley additive explanations; SVM: support vector machine.
LIME
LIME is used to explain how features contribute to model predictions at the local level. This facilitates interpretation of complex decisions and reveals differences in focus between algorithms for each preeclampsia risk category.
In Figure 8, LIME analysis reveals the differences in focus of each algorithm. It highlighted model-specific local decision patterns, reinforcing the complementary role of local explanations alongside global interpretability methods. Although blood pressure variables were consistently among the most influential features across models, their relative importance varied depending on the algorithm and explainability method.
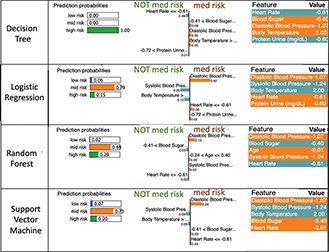
Fig. 8. LIME visualization for explaining the model’s prediction of preeclampsia risk. BS: blood sugar; BT: body temperature; DBP: diastolic blood pressure; HR: heart rate; LIME: local interpretable model-agnostic explanations; SBP: systolic blood pressure.
Visualization Permutation Feature Importance
PFI visualization is used to assess the contribution of each feature to model performance. A feature’s importance is determined by how much accuracy decreases when its value is randomized. The largest decrease indicates the feature’s most significant influence on preeclampsia risk prediction.
PFI (Figure 9) shows feature sensitivity differed across models. While blood pressure variables frequently contributed to performance changes, other features, such as blood glucose and body temperature, played model-dependent roles.
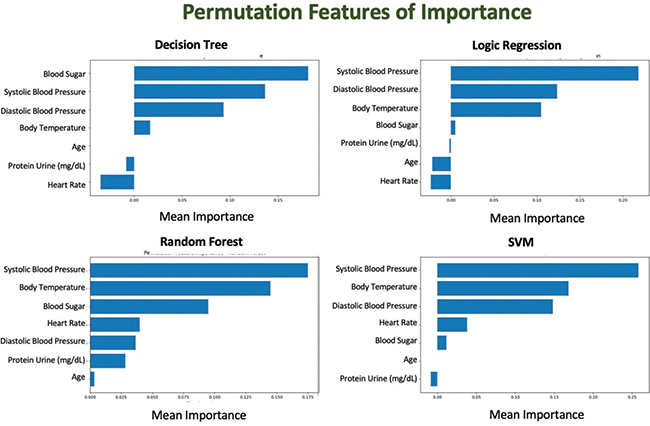
Fig. 9. Permutation Feature Importance (PFI) visualization: Showing the decrease in model performance when each feature is randomly permuted. SVM: support vector machine.
Discussion
The consistent model performance across multiple evaluation metrics, including accuracy, recall, the F1 score, and the AUC, indicates stable internal performance. A key strength lies in the focused evaluation of the medium-risk group, which is the most clinically relevant and challenging to classify. Furthermore, integrating multi-method XAI (SHAP, LIME, and PFI) enhances the interpretability of the model, with comprehensive XAI visualizations facilitating an understanding of the prediction mechanisms.
Notably, the relative importance of features varies depending on the explainability method and learning algorithm used, reflecting the fundamental differences between global interpretability frameworks, such as SHAP and PFI, and local interpretability frameworks, such as LIME. These variations suggest that feature dominance depends on the model and method rather than being absolute. Therefore, XAI findings should be viewed as different perspectives on model behavior rather than definitive rankings of clinical importance.
From a clinical screening perspective, misclassification, particularly false-negative predictions in the medium-risk group, might delay closer monitoring or early intervention. This highlights the importance of cautious interpretation and reinforces that the proposed models are intended to support early risk screening rather than definitive diagnosis.
The RF and SVM algorithms deliver the most accurate predictions. They achieve an accuracy rate of 78%, a precision rate of 0.77, and an F1-score of 0.77. The main advantage of this study lies in the application of the multi-method XAI approach, which combines SHAP, LIME, and PFI methods to improve model transparency and interpretability. This improves the technical transparency of model behavior and feature contributions. However, because no formal usability testing or expert evaluation involving clinicians was conducted, interpretability claims are limited to methodological assessment rather than validated clinical usability.
Due to differences in datasets, outcome definitions, and validation strategies, direct comparisons with previous studies should be interpreted cautiously. Nevertheless, the proposed models demonstrate balanced internal performance within the context of the available data. Studies (22) and (34) utilized one algorithm, which achieves accuracies of 79.5 and 65.22%, respectively. Studies (31) and (16) emphasized the role of AI in improving accuracy; the best model (SGB) achieved 97.3% accuracy but was not highly interpretable. Therefore, this study’s advantage lies not only in the stability and balance of the model’s performance but in its ability to identify the most influential variables (e.g., blood pressure and blood glucose levels) in predicting preeclampsia risk.
All the predictors used in this study were collected at a single time point using a cross-sectional design. While this approach is suitable for creating a baseline screening model, it is not adequate to support long-term clinical inference or analyzing disease progression. Future research should therefore incorporate longitudinal or trend-based clinical data in order to capture the dynamic physiological changes that occur during pregnancy. This could enhance risk stratification and improve the predictive performance of preeclampsia screening models.
Although RF and SVM achieved the highest overall accuracy (78%) and stable classification across all risk groups, this remains a moderate level for clinical application. False negative predictions within the medium-risk group, in particular, may delay early intervention and allow progression to high-risk preeclampsia without timely monitoring. From a clinical perspective, the balanced performance observed across risk categories, especially within the medium-risk group, suggests that the proposed models could serve as screening aids rather than definitive diagnostic systems. Future work should involve testing the model’s transportability across diverse populations.
Conclusion
Four ML algorithms, which were utilized, are logistic regression, decision tree, SVM, and RF. The evaluation used the multi-method XAI approach, which combines SHAP, LIME, and PFI. The results showed that RF and SVM provided the best performance. They achieved 78% of accuracy and made consistent predictions across all risk categories. Multi-method XAI analysis indicated that blood pressure variables frequently appeared among influential features, although their relative importance varied across models and explainability methods. The multi-method XAI approach not only improves accuracy but also provides transparency and interpretability by enhancing methodological transparency and supporting model interpretability.
These findings might support future decision-making based on screening by providing transparent insight into model behavior. However, further clinical validation is required before real-world implementation. Therefore, the proposed model should be interpreted as a tool to support early risk screening and assist healthcare professionals in identifying potential preeclampsia risk, rather than as a diagnostic system or a substitute for clinical judgment.
Ethical Approval
Ethical clearance for this study was obtained from the Anak Air Community Health Centre (Puskesmas Anak Air) in the Koto Tangah district of Padang city. A written informed consent was obtained from all participants prior to data collection. To ensure confidentiality and compliance with ethical research standards, all personal identifiers were removed, and the data were fully anonymized before analysis.
Contributions
Sari Puspita supervised the research, administered the project, provided resources, and critically revised the manuscript. Gusrino Yanto conceptualized the study, collected and processed the data, performed machine learning analysis, implemented the Multi-Methode XAI, and drafted the original manuscript. Rifa Turaina and Nency Extise Putri contributed to manuscript refinement and critical review.
Data Availability Statement (DAS), Data Sharing, Reproducibility, and Data Repositories
The data that support the findings of this study are available from the corresponding author upon reasonable request.
Application of AI-Generated Text or Related Technology
AI-assisted tools, including language refinement and grammar correction software, were used for improving linguistic clarity and formatting of the manuscript. No part of the analytical work, data-set processing, or result generation was performed using AI-generated content.
Ethics/IRB Statement
The study used retrospective secondary data collected in 2024 from the Anak Air community health center. The dataset was fully anonymized prior to being provided to the authors, and no identifiable personal information was accessed during the analysis. According to institutional research guidelines, retrospective analysis of anonymized secondary data does not require formal IRB approval. Therefore, an ethics approval number is not applicable. Informed consent was waived due to institutional data protection policies, but may be made available from the corresponding author upon reasonable request. The study was not prospectively registered because it was not a prospective clinical study.
References
- Brownfoot F, Rolnik DL. Prevention of preeclampsia. Best Pract Res Clin Obstet Gynaecol. 2024;93:102481. https://doi.org/10.1016/j.bpobgyn.2024.102481
- Erez O, Romero R, Jung E, Chaemsaithong P, Bosco M, Suksai M, et al. Preeclampsia and eclampsia: the conceptual evolution of a syndrome. Am J Obstet Gynecol. 2022;226:S786–803. https://doi.org/10.1016/j.ajog.2021.12.001
- Aziz F, Khan MF, Moiz A. Gestational diabetes mellitus, hypertension, and dyslipidemia as the risk factors of preeclampsia. Sci Rep. 2024;14(1):6182. https://doi.org/10.1038/s41598-024-56790-z
- Yang S, Zhou W, Dimitriadis E, Menkhorst E. Maternal blood lipoprotein cholesterol prior to and at the time of diagnosis of preeclampsia: a systematic review. Am J Obstet Gynecol MFM. 2025;7(5):101654. https://doi.org/10.1016/j.ajogmf.2025.101654
- Auger N, Ayoub A, Bilodeau-Bertrand M, Lafleur N, Wei SQ. Ethnocultural status and risk of preeclampsia in a Canadian setting. Pregnancy Hypertens. 2025;39:101202. https://doi.org/10.1016/j.preghy.2025.101202
- Ives CW, Sinkey R, Rajapreyar I, Tita ATN, Oparil S. Preeclampsia—pathophysiology and clinical presentations: JACC state-of-the-art review. J Am Coll Cardiol. 2020;76:1690–702. https://doi.org/10.1016/j.jacc.2020.08.014
- Bergman L, Hannsberger D, Schell S, Imberg H, Langenegger E, Moodley A, et al. Cerebral infarcts, edema, hypoperfusion, and vasospasm in preeclampsia and eclampsia. Am J Obstet Gynecol. 2025;232(6):550.e1–14. https://doi.org/10.1016/j.ajog.2024.10.034
- Lee ST, Lee YL, Chen YC, Lin W, Wu CI, Lin CK. Arteriovenous malformation-related headache and seizures in pregnancy masquerading as eclampsia: a case report. Taiwan J Obstet Gynecol. 2024;63(4):552–6. https://doi.org/10.1016/j.tjog.2023.09.026
- Shinde S, Yelverton CA, Yussuf M, Nurhussien L, Wang D, Fawzi WW. Effects of vitamin and multiple micronutrient supplementation for pregnant and/or lactating women on maternal and infant nutritional status in low- and middle-income countries: a systematic review and meta-analysis. Adv Nutr. 2025;16(12):100487. https://doi.org/10.1016/j.advnut.2025.100487
- Liu Z, Zhang Z, Yang H, Wang G, Xu Z. An innovative model fusion algorithm to improve the recall rate of peer-to-peer lending default customers. Intell Syst Appl. 2023;20:200272. https://doi.org/10.1016/j.iswa.2023.200272
- Liu Q, Li J, Li Y. Safety of pertussis vaccination in pregnancy and effectiveness in infants: a Danish national cohort study 2019–2023. Clin Microbiol Infect. 2025;31(6):995–1002. https://doi.org/10.1016/j.cmi.2025.03.014
- Macdonald TM, Walker SP, Hannan NJ, Tong S, Uhevaha T, Kaitu’u-Lino J. Clinical tools and biomarkers to predict preeclampsia. EBioMedicine. 2022;75:103780. https://doi.org/10.1016/j.ebiom.2021.103780
- Talukdar D, Sarkar M, Ahrodia T, Kumar S, De D, Nath S, et al. Previse preterm birth in early pregnancy through vaginal microbiome signatures using metagenomics and dipstick assays. iScience. 2024;27(11):111238. https://doi.org/10.1016/j.isci.2024.111238
- Brown MA, Magee LA, Kenny LC, Karumanchi SA, McCarthy FP, Saito S, et al. Hypertensive disorders of pregnancy: ISSHP classification, diagnosis, and management recommendations for international practice. Hypertension. 2018;72:24–43. https://doi.org/10.1161/HYPERTENSIONAHA.117.10803
- Ansbacher-Feldman Z, Syngelaki A, Meiri H, Cirkin R, Nicolaides KH, Louzoun Y. Machine-learning-based prediction of pre-eclampsia using first-trimester maternal characteristics and biomarkers. Ultrasound Obstet Gynecol. 2022;60(6):739–45. https://doi.org/10.1002/uog.26105
- Jhee JH, Lee S, Park Y, Lee SE, Kim YA, Kang SW, et al. Prediction model development of late-onset preeclampsia using machine learning-based methods. PLoS One. 2019;14(8):e0221202. https://doi.org/10.1371/journal.pone.0221202
- Liu M, Yang X, Chen G, Ding Y, Shi M, Sun L, et al. Development of a prediction model on preeclampsia using machine learning-based method: a retrospective cohort study in China. Front Physiol. 2022;13:896969. https://doi.org/10.3389/fphys.2022.896969
- Kovacheva VP, Eberhard BW, Cohen RY, Maher M, Saxena R, Gray KJ. Preeclampsia prediction using machine learning and polygenic risk scores from clinical and genetic risk factors in early and late pregnancies. Hypertension. 2024;81(2):264–72. https://doi.org/10.1161/HYPERTENSIONAHA.123.21053
- Tiruneh SA, Rolnik DL, Teede HJ, Enticott J. Prediction of pre-eclampsia with machine learning approaches: leveraging important information from routinely collected data. Int J Med Inform. 2024;192:105645. https://doi.org/10.1016/j.ijmedinf.2024.105645
- Wang L, Ma Y, Bi W, Meng C, Liang X, Wu H, et al. An early screening model for preeclampsia: utilizing zero-cost maternal predictors exclusively. Hypertens Res. 2024;47(4):1051–62. https://doi.org/10.1038/s41440-023-01573-8
- Khalil A, Bellesia G, Norton ME, Jacobsson B, Haeri S, Egbert M, et al. The role of cell-free DNA biomarkers and patient data in the early prediction of preeclampsia: an artificial intelligence model. Am J Obstet Gynecol. 2024;231(5):554.e1–18. https://doi.org/10.1016/j.ajog.2024.02.299
- Wesson JL, Smith N. A machine learning model to predict preeclampsia in pregnant women. Proc Comput Sci. 2024;239:1645–52. https://doi.org/10.1016/j.procs.2024.06.341
- Kota S. Kota Padang Dalam Angka 2024 [Internet]. Bps.go.id. Badan Pusat Statistik Kota Padang; 2024 [cited 2025 Nov 20]. Available from: https://padangkota.bps.go.id/id/publication/2024/02/28/c4991c8e8aeffe085e50de1e/kota-padang-dalam-angka-2024.html
- DINAS KESEHATAN KOTA PADANG [Internet]. Padang.go.id. 2022 [cited 2025 Aug 21]. Available from: https://dinkes.padang.go.id/profil-kesehatan-dinkes
- DINAS KESEHATAN KOTA PADANG [Internet]. Padang.go.id. 2024 [cited 2025 Aug 21]. Available from: https://dinkes.padang.go.id/laporan-tahunan-dinkes
- Ye H. Artificial intelligence combined with computed tomography or X-ray radiography: potential solution for opportunistic screening for osteoporosis. Eur J Radiol Artific Intell. 2025;3:100036. https://doi.org/10.1016/j.ejrai.2025.100036
- Kyriakopoulos N, Kim E, Hultink EJ, Santema S. The impact of design thinking and artificial intelligence capabilities on performance: the role of new product development decision-making agility. J Bus Res. 2025;200:115633. https://doi.org/10.1016/j.jbusres.2025.115633
- García-Torres M, Saucedo F, Divina F, Gómez-Guerrero S. RFMSU: a multivariate symmetrical uncertainty-based random forest. Pattern Recognit. 2026;169:111939. https://doi.org/10.1016/j.patcog.2025.111939
- Malik V, Agrawal N, Prasad S, Talwar S, Khatuja R, Jain S, et al. Prediction of preeclampsia using machine learning: a systematic review. Cureus. 2024;16(12):e76095. https://doi.org/10.7759/cureus.76095
- Chen X, Chen H, Nan S, Kong X, Duan H, Zhu H. Dealing with missing, imbalanced, and sparse features during the development of a prediction model for sudden death using emergency medicine data: machine learning approach. JMIR Med Inform. 2023;11:e38590. https://doi.org/10.2196/38590
- Layton AT. Artificial intelligence and machine learning in preeclampsia. Arterioscler Thromb Vasc Biol. 2025;45(2):165–71. https://doi.org/10.1161/ATVBAHA.124.321673
- Wahyuningsih T, Manongga D, Sembiring I, Wijono S. Comparison of effectiveness of logistic regression, naive bayes, and random forest algorithms in predicting student arguments. Proc Comput Sci. 2024;234:349–56. https://doi.org/10.1016/j.procs.2024.03.014
- Hassanpouri Baesmat K, Shokoohi F, Farrokhi Z. SP-RF-ARIMA: a sparse random forest and ARIMA hybrid model for electric load forecasting. Glob Energy Interconnect. 2025;8(3):486–96. https://doi.org/10.1016/j.gloei.2025.04.003
- Harizahayu H, Friendly F, Purwo Seputro B, Benar B, Hermanto K. Predictive modeling of preeclampsia risk using random forest algorithm within a machine learning framework. J Comput Netw Architect High Perform Comput. 2024;6(4):1843–50. https://doi.org/10.47709/cnahpc.v6i4.4779
- Yanto G, Puspita S. A random forest algorithm for high-risk pregnancies prediction based on explainable artificial intelligence (XAI). Commun Math Biol Neurosci. 2025;2025:126. https://doi.org/10.28919/cmbn/9536
- Supsermpol P, Huynh VN, Thajchayapong S, Chiadamrong N. Predicting financial performance for listed companies in Thailand during the transition period: a class-based approach using logistic regression and random forest algorithm. J Open Innov Technol Market Complex. 2023;9(3):100130. https://doi.org/10.1016/j.joitmc.2023.100130
- Belsti Y, Moran L, Du L, Mousa A, De Silva K, Enticott J, et al. Comparison of machine learning and conventional logistic regression-based prediction models for gestational diabetes in an ethnically diverse population; the Monash GDM Machine learning model. Int J Med Inform. 2023;179:105228. https://doi.org/10.1016/j.ijmedinf.2023.105228
- Lakshmi BN, Indumathi TS, Ravi N. A study on C.5 decision tree classification algorithm for risk predictions during pregnancy. Proced Technol. 2016;24:1542–9. https://doi.org/10.1016/j.protcy.2016.05.128
- Zhao R, Hong L, Ji H, Zhang Q, Zhang S, Li Q, et al. Decision tree based parameter identification and state estimation: application to reactor operation Digital Twin. Nucl Eng Technol. 2025;57:103527. https://doi.org/10.1016/j.net.2025.103527
- Gao Y, Wang Y, Tian L, Hong X, Xue C, Li D. Evolving adaptive and interpretable decision trees for cooperative submarine search. Def Technol. 2025;48:83–94. https://doi.org/10.1016/j.dt.2025.02.007
- Jiang N, Li P, Feng Z. Detecting tropical freshly-opened swidden fields using a combined algorithm of continuous change detection and support vector machine. Int J Appl Earth Observ Geoinform. 2025;136:104403. https://doi.org/10.1016/j.jag.2025.104403
- Shetty NP, Shetty J, Hegde V, Dharne SD, Kv M. A machine learning-based clinical decision support system for effective stratification of gestational diabetes mellitus and management through Ayurveda. J Ayurveda Integr Med. 2024;15(6):101051. https://doi.org/10.1016/j.jaim.2024.101051
- Piazza M, Spinelli A, Maggioni F, Bedoni M, Messina E. A robust support vector machine approach for Raman data classification. Decis Analyt J. 2025;16:100595. https://doi.org/10.1016/j.dajour.2025.100595
- Raghunath MP, Deshmukh S, Chaudhari P, Bangare SL, Kasat K, Awasthy M, et al. PCA and PSO based optimized support vector machine for efficient intrusion detection in internet of things. Measur Sens. 2025;37:101806. https://doi.org/10.1016/j.measen.2024.101806
- Mohamad Deros SN, Naim MR, Din NM. Explainable artificial intelligence (XAI) to interpret the contributing factors of land subsidence susceptibility prediction model. Total Environ Adv. 2025;15:200129. https://doi.org/10.1016/j.teadva.2025.200129
- Nimmy SF, Hussain OK, Chakrabortty RK, Saha S. Explainable artificial intelligence (XAI) in glaucoma assessment: advancing the frontiers of machine learning algorithms. Knowl Based Syst. 2025;316:113333. https://doi.org/10.1016/j.knosys.2025.113333
- Umaa Mahesswari G, Uma Maheswari P. SmartScanPCOS: a feature-driven approach to cutting-edge prediction of polycystic ovary syndrome using machine learning and explainable artificial intelligence. Heliyon. 2024;10(20):e39205. https://doi.org/10.1016/j.heliyon.2024.e39205
- Han J, Guzman JA, Chu ML. Prediction of gully erosion susceptibility through the lens of the SHapley Additive exPlanations (SHAP) method using a stacking ensemble model. J Environ Manage. 2025;383:125478. https://doi.org/10.1016/j.jenvman.2025.125478
- Mehdiyev N, Majlatow M, Fettke P. Integrating permutation feature importance with conformal prediction for robust explainable artificial intelligence in predictive process monitoring. Eng Appl Artif Intell. 2025;149:110363. https://doi.org/10.1016/j.engappai.2025.110363
- Ramirez SG, Hales RC, Williams GP, Jones NL. Extending SC-PDSI-PM with neural network regression using GLDAS data and permutation feature importance. Environ Modell Softw. 2022;157:105475. https://doi.org/10.1016/j.envsoft.2022.105475
- Hassan SU, Abdulkadir SJ, Zahid MSM, Al-Selwi SM. Local interpretable model-agnostic explanation approach for medical imaging analysis: a systematic literature review. Comput Biol Med. 2025;185:109569. https://doi.org/10.1016/j.compbiomed.2024.109569
- Ali M. Classification of imbalanced travel mode choice dataset with SMOTE and prediction using interpretable machine learning. Sustain Fut. 2025;10:101119. https://doi.org/10.1016/j.sftr.2025.101119
- Aruleba I, Sun Y. Enhanced credit risk prediction using deep learning and SMOTE-ENN resampling. Mach Learn Appl. 2025;21:100692. https://doi.org/10.1016/j.mlwa.2025.100692
- Yusoff M, Mahmud Y, Azmi PAR, Sallehud-din MTM. The improvement of SMOTE-ENN-XGBoost through Yeo Johnson strategy on dissolved gas analysis dataset. Ener Rep. 2025;13:6281–90. https://doi.org/10.1016/j.egyr.2025.05.013
- Kohan AA, Mirshahvalad SA, Hinzpeter R, Kulanthaivelu R, Avery L, Ortega C, et al. External validation of a CT-based radiogenomics model for the detection of EGFR mutation in NSCLC and the impact of prevalence in model building by using Synthetic Minority Over Sampling (SMOTE): lessons learned. Acad Radiol. 2025;32(9):5576–84. https://doi.org/10.1016/j.acra.2025.04.064
- Shamshuzzoha M, Audry TTB, Alam MJ, Bhuiyan ZA, Motaharul Islam M, Hassan MM. A novel framework for seasonal affective disorder detection: comprehensive machine learning analysis using multimodal social media data and SMOTE. Acta Psychol (Amst). 2025;256:105005. https://doi.org/10.1016/j.actpsy.2025.105005
- Dash D, Kumar M, Patra S, Kumar A, Ganguly A. Healthcare fraud detection using an integrated ML approach with SMOTE. Proc Comput Sci. 2025;258:800–10. https://doi.org/10.1016/j.procs.2025.04.312
- Xu H, Li H, Fan Y, Wang Y, Li Z, Zhou L, et al. Analysis of factors influencing chemotherapy-induced peripheral neuropathy in breast cancer patients using a random forest model. Breast. 2025;81:104457. https://doi.org/10.1016/j.breast.2025.104457
- Hapsari GI, Munadi R, Erfianto B, Irawati ID. Feature selection using Pearson correlation for ultra-wideband ranging classification. J RESTI. 2025;9(2):209–17. https://doi.org/10.29207/resti.v9i2.6281
- Li W, Chen S, Lin L, Chen L. Random-forest-based task pricing model and task-accomplished model for crowdsourced emergency information acquisition. Syst Soft Comput. 2025;7:200235. https://doi.org/10.1016/j.sasc.2025.200235
- Mehaba N, Schrade S, Eggerschwiler L, Dohme-Meier F, Schlegel P. Accuracy and precision in DM intake prediction models for lactating dairy cows. Animal. 2025;19(7):101535. https://doi.org/10.1016/j.animal.2025.101535
- Fukuda S, Yamamoto N, Tomita Y, Matsumoto T, Shinohara T, Ohno T, et al. Development and validation of clinical prediction model for functional independence measure following stroke rehabilitation. J Stroke Cerebrovasc Dis. 2025;34(2):108185. https://doi.org/10.1016/j.jstrokecerebrovasdis.2024.108185
- Singh MP, Bisht N, Choudhary M, Goswami A, Tagore NK. A web-based supervised machine learning model for air quality index and respiratory care prediction. Proc Comput Sci. 2025;258:1747–56. https://doi.org/10.1016/j.procs.2025.04.426
- Zanchi M, Zapperi S, Bocchi S, Drofa O, Davolio S, La Porta CAM. Improving localized weather predictions for precision agriculture: a time-series mixer approach for hazardous event detection. Environ Modell Softw. 2025;191:106509. https://doi.org/10.1016/j.envsoft.2025.106509
- Kumar V, prabha C, Gupta D, Juneja S, Kumari S, Nauman A. Multi-model machine learning framework for lung cancer risk prediction: a comparative analysis of nine classifiers with hybrid and ensemble approaches using behavioral and hematological parameters. SLAS Technol. 2025;33:100314. https://doi.org/10.1016/j.slast.2025.100314
- Soladoye AA, Aderinto N, Omodunbi BA, Esan AO, Adeyanju IA, Olawade DB. Enhancing Alzheimer’s disease prediction using random forest: a novel framework combining backward feature elimination and ant colony optimization. Curr Res Transl Med. 2025;73(4):103526. https://doi.org/10.1016/j.retram.2025.103526
- Zuege CV, Stefenon SF, Yamaguchi CK, Mariani VC, Gonzalez GV, dos Santos Coelho L. Wind speed forecasting approach using conformal prediction and feature importance selection. Int J Electr Power Ener Syst. 2025;168:110700. https://doi.org/10.1016/j.ijepes.2025.110700
- Abekoon T, Sajindra H, Rathnayake N, Ekanayake IU, Jayakody A, Rathnayake U. A novel application with explainable machine learning (SHAP and LIME) to predict soil N, P, and K nutrient content in cabbage cultivation. Smart Agric Technol. 2025;11:100879. https://doi.org/10.1016/j.atech.2025.100879
- Shahzad MF, Xu S, Lim WM, Yang X, Khan QR. Artificial intelligence and social media on academic performance and mental well-being: student perceptions of positive impact in the age of smart learning. Heliyon. 2024;10(8):e29523. https://doi.org/10.1016/j.heliyon.2024.e29523
- Velasquez JD. TechMiner: analysis of bibliographic datasets using python. SoftwareX. 2023;23:101457. https://doi.org/10.1016/j.softx.2023.101457
- Horsburgh JS, Black S, Castronova A, Dash PK. Advancing open and reproducible water data science by integrating data analytics with an online data repository. Environ Modell Softw. 2025;188:106422. https://doi.org/10.1016/j.envsoft.2025.106422
Copyright Ownership: This is an open-access article distributed in accordance with the Creative Commons Attribution Non-Commercial (CC BY-NC 4.0) license, which permits others to distribute, adapt, enhance this work non-commercially, and license their derivative works on different terms, provided the original work is properly cited and the use is non-commercial. See http://creativecommons.org/licenses/by-nc/4.0. The authors of this article own the copyright.